Proteomics data analysis: heart
Lieven Clement
statOmics, Ghent University (https://statomics.github.io)
1 Background
Researchers have assessed the proteome in different regions of the heart for 3 patients (identifiers 3, 4, and 8). For each patient they sampled the left atrium (LA), right atrium (RA), left ventricle (LV) and the right ventricle (RV). The data are a small subset of the public dataset PXD006675 on PRIDE.
Suppose that researchers are mainly interested in comparing the ventricular to the atrial proteome. Particularly, they would like to compare the left atrium to the left ventricle, the right atrium to the right ventricle, the average ventricular vs atrial proteome and if ventricular vs atrial proteome shifts differ between left and right heart region.
2 Data
We first import the peptides.txt file. This is the file that contains your peptide-level intensities. For a MaxQuant search [6], this peptides.txt file can be found by default in the “path_to_raw_files/combined/txt/” folder from the MaxQuant output, with “path_to_raw_files” the folder where raw files were saved. In this tutorial, we will use a MaxQuant peptides file from MaxQuant that can be found in the data tree of the SGA2020 github repository https://github.com/statOmics/SGA2020/tree/data/quantification/heart .
To import the data we use the QFeatures package.
We generate the object peptideRawFile with the path to the
peptideRaws.txt file. Using the grepEcols function, we find
the columns that contain the expression data of the peptideRaws in the
peptideRaws.txt file.
library(tidyverse)
library(limma)
library(QFeatures)
library(msqrob2)
library(plotly)
peptidesFile <- "https://raw.githubusercontent.com/statOmics/PDA21/data/quantification/heart/peptides.txt"
ecols <- grep("Intensity\\.", names(read.delim(peptidesFile)))
pe <- readQFeatures(
assayData = read.delim(peptidesFile),
fnames = 1,
quantCols = ecols,
name = "peptideRaw")
pe## An instance of class QFeatures containing 1 assays:
## [1] peptideRaw: SummarizedExperiment with 31319 rows and 12 columns## class: SummarizedExperiment
## dim: 31319 12
## metadata(0):
## assays(1): ''
## rownames(31319): AAAAAAAAAK AAAAAAAAEQQSSNGPVK ... YYTPVPCESATAK
## YYTYLIMNK
## rowData names(91): Sequence N.term.cleavage.window ...
## Oxidation..M..site.IDs MS.MS.Count
## colnames(12): Intensity.LA3 Intensity.LA4 ... Intensity.RV4
## Intensity.RV8
## colData names(0):We will make use from data wrangling functionalities from the tidyverse package. The %>% operator allows us to pipe the output of one function to the next function.
colData(pe)$location <- substr(
colnames(pe[["peptideRaw"]]),
11,
11) %>%
unlist %>%
as.factor
colData(pe)$tissue <- substr(
colnames(pe[["peptideRaw"]]),
12,
12) %>%
unlist %>%
as.factor
colData(pe)$patient <- substr(
colnames(pe[["peptideRaw"]]),
13,
13) %>%
unlist %>%
as.factorWe calculate how many non zero intensities we have per peptide and this will be useful for filtering.
Peptides with zero intensities are missing peptides and should be
represent with a NA value rather than 0.
2.1 Data exploration
63% of all peptide intensities are missing and for some peptides we do not even measure a signal in any sample. The missingness is similar across samples.
3 Preprocessing
This section preforms standard preprocessing for the peptide data. This include log transformation, filtering and summarisation of the data.
3.1 Log transform the data
pe <- logTransform(pe, base = 2, i = "peptideRaw", name = "peptideLog")
limma::plotDensities(assay(pe[["peptideLog"]]))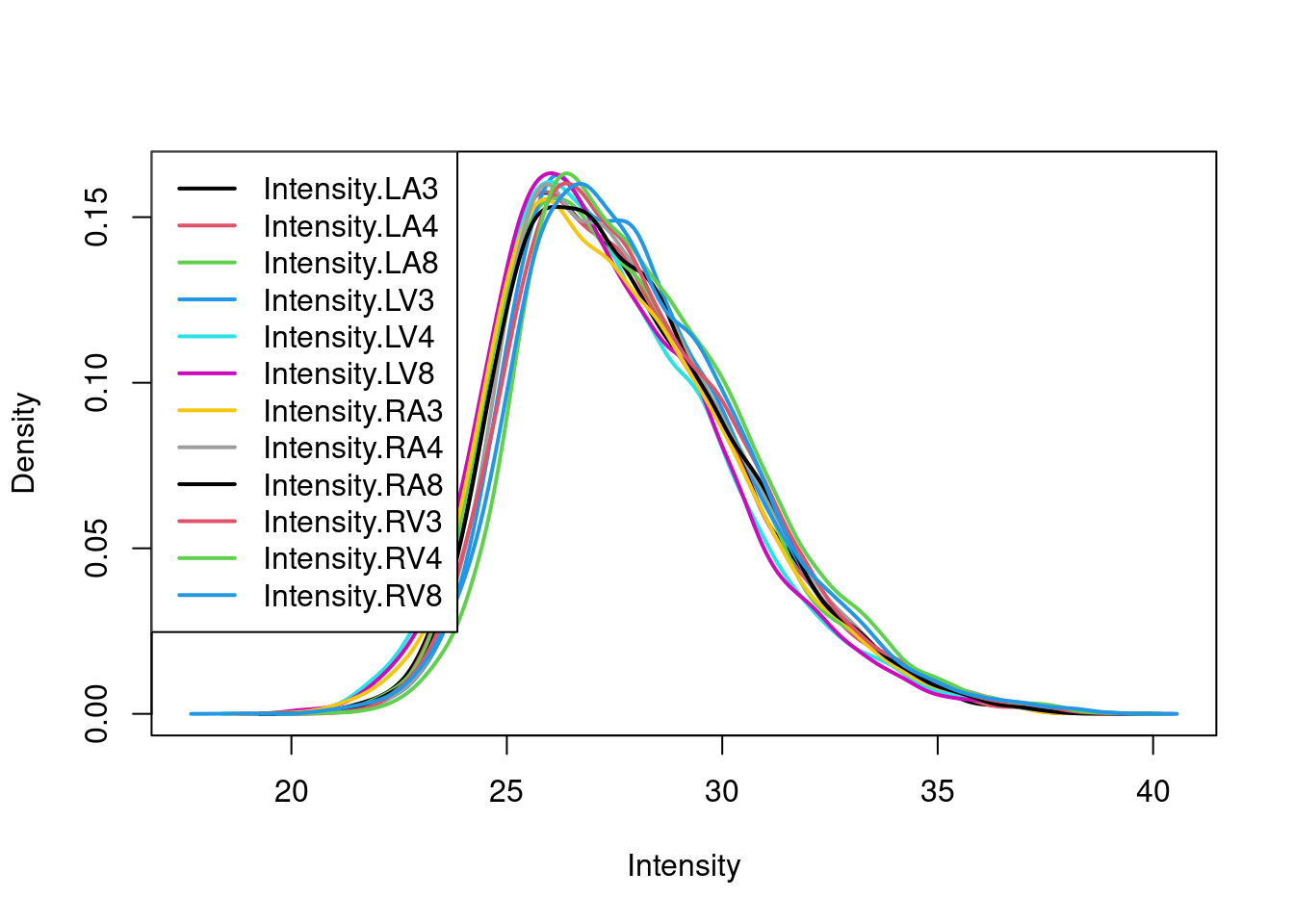
3.2 Filtering
3.2.1 Handling overlapping protein groups
In our approach a peptide can map to multiple proteins, as long as there is none of these proteins present in a smaller subgroup.
pe <- filterFeatures(pe, ~ Proteins %in% smallestUniqueGroups(rowData(pe[["peptideLog"]])$Proteins))## 'Proteins' found in 2 out of 2 assay(s)3.2.2 Remove reverse sequences (decoys) and contaminants
We now remove the contaminants, peptides that map to decoy sequences, and proteins which were only identified by peptides with modifications.
First look to the names of the variables for the peptide features
## [1] "Sequence" "N.term.cleavage.window"
## [3] "C.term.cleavage.window" "Amino.acid.before"
## [5] "First.amino.acid" "Second.amino.acid"
## [7] "Second.last.amino.acid" "Last.amino.acid"
## [9] "Amino.acid.after" "A.Count"
## [11] "R.Count" "N.Count"
## [13] "D.Count" "C.Count"
## [15] "Q.Count" "E.Count"
## [17] "G.Count" "H.Count"
## [19] "I.Count" "L.Count"
## [21] "K.Count" "M.Count"
## [23] "F.Count" "P.Count"
## [25] "S.Count" "T.Count"
## [27] "W.Count" "Y.Count"
## [29] "V.Count" "U.Count"
## [31] "O.Count" "Length"
## [33] "Missed.cleavages" "Mass"
## [35] "Proteins" "Leading.razor.protein"
## [37] "Start.position" "End.position"
## [39] "Gene.names" "Protein.names"
## [41] "Unique..Groups." "Unique..Proteins."
## [43] "Charges" "PEP"
## [45] "Score" "Identification.type.LA3"
## [47] "Identification.type.LA4" "Identification.type.LA8"
## [49] "Identification.type.LV3" "Identification.type.LV4"
## [51] "Identification.type.LV8" "Identification.type.RA3"
## [53] "Identification.type.RA4" "Identification.type.RA8"
## [55] "Identification.type.RV3" "Identification.type.RV4"
## [57] "Identification.type.RV8" "Fraction.Average"
## [59] "Fraction.Std..Dev." "Fraction.1"
## [61] "Fraction.2" "Fraction.3"
## [63] "Fraction.4" "Fraction.5"
## [65] "Fraction.6" "Fraction.7"
## [67] "Fraction.8" "Fraction.100"
## [69] "Experiment.LA3" "Experiment.LA4"
## [71] "Experiment.LA8" "Experiment.LV3"
## [73] "Experiment.LV4" "Experiment.LV8"
## [75] "Experiment.RA3" "Experiment.RA4"
## [77] "Experiment.RA8" "Experiment.RV3"
## [79] "Experiment.RV4" "Experiment.RV8"
## [81] "Intensity" "Reverse"
## [83] "Potential.contaminant" "id"
## [85] "Protein.group.IDs" "Mod..peptide.IDs"
## [87] "Evidence.IDs" "MS.MS.IDs"
## [89] "Best.MS.MS" "Oxidation..M..site.IDs"
## [91] "MS.MS.Count" "nNonZero"No information on decoys.
## 'Potential.contaminant' found in 2 out of 2 assay(s)3.3 Normalize the data
3.4 Explore normalized data
After normalisation the density curves for all samples are comparable.
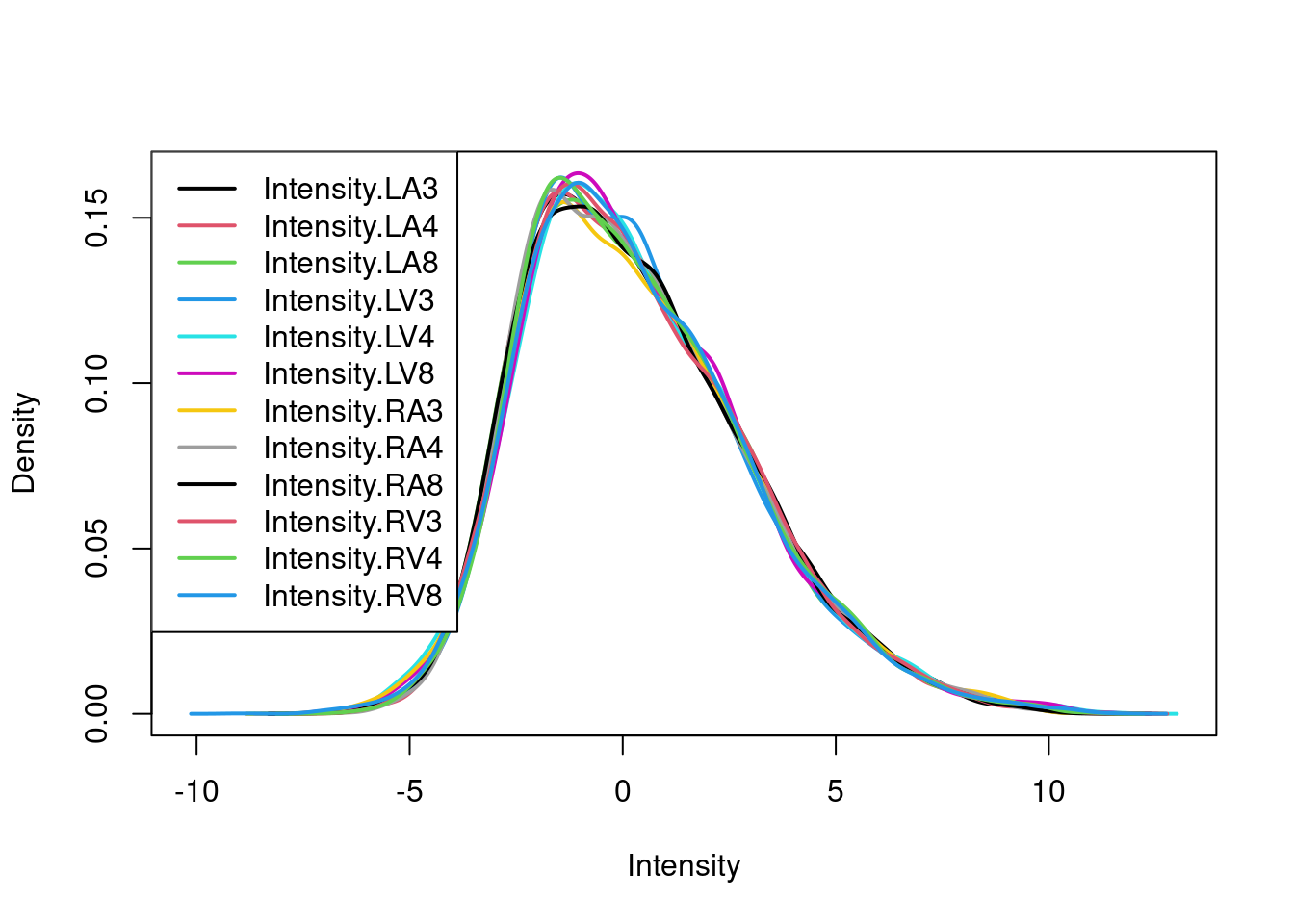
This is more clearly seen is a boxplot.
boxplot(assay(pe[["peptideNorm"]]), col = palette()[-1],
main = "Peptide distribtutions after normalisation", ylab = "intensity")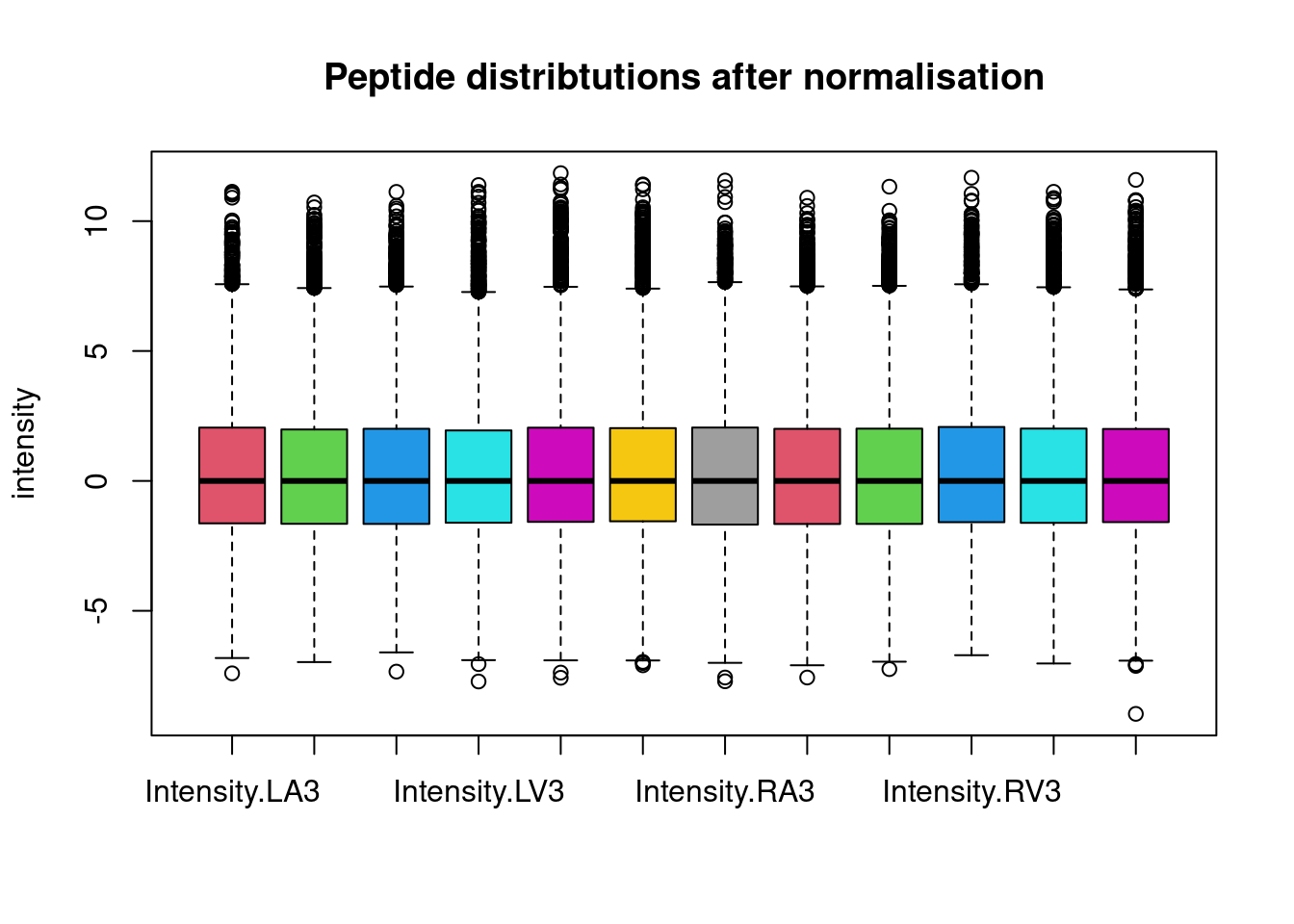
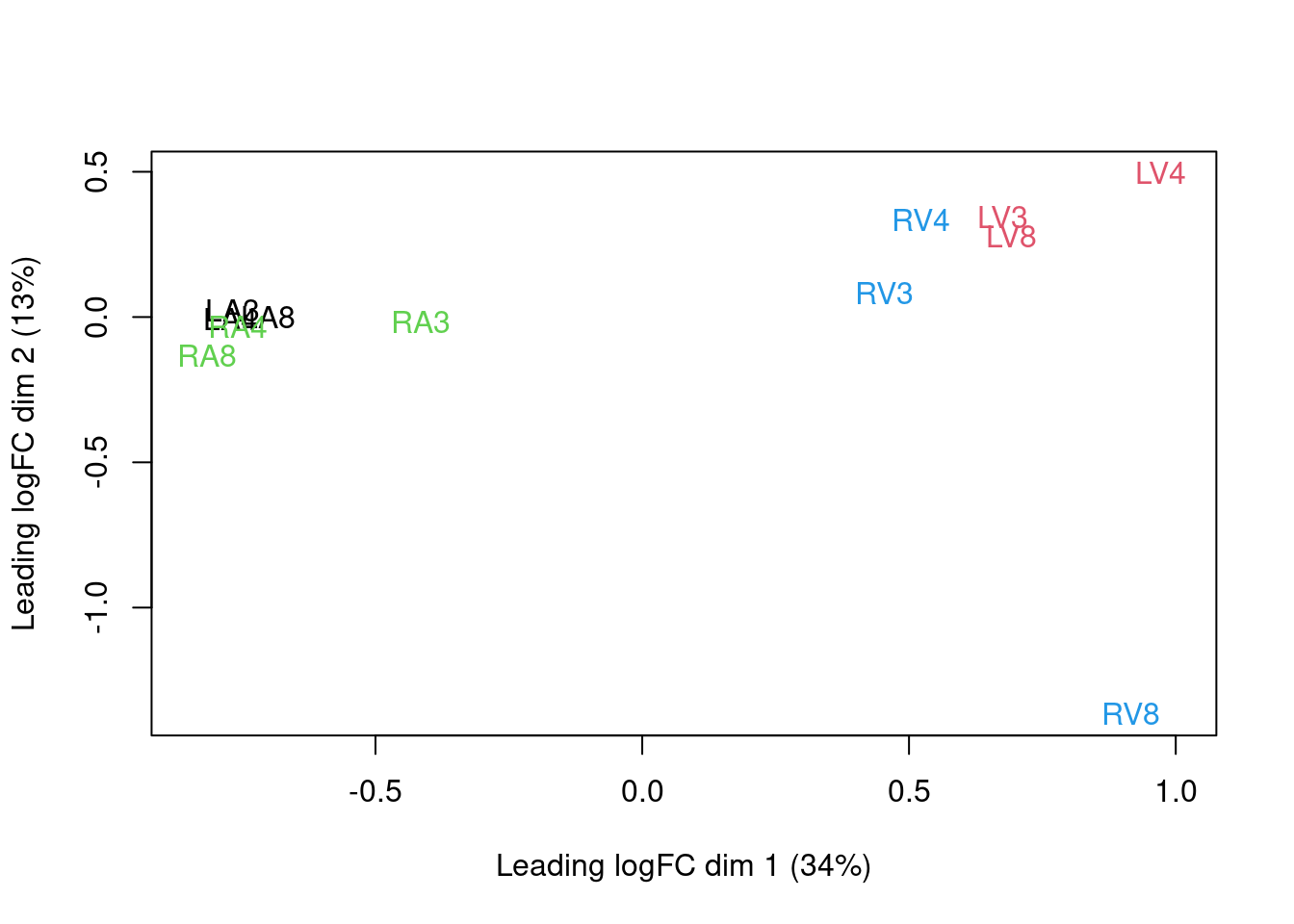
We can visualize our data using a Multi Dimensional Scaling plot, eg.
as provided by the limma package.
limma::plotMDS(assay(pe[["peptideNorm"]]),
col = colData(pe)$location:colData(pe)$tissue %>%
as.numeric,
labels = colData(pe) %>%
rownames %>%
substr(start = 11, stop = 13)
)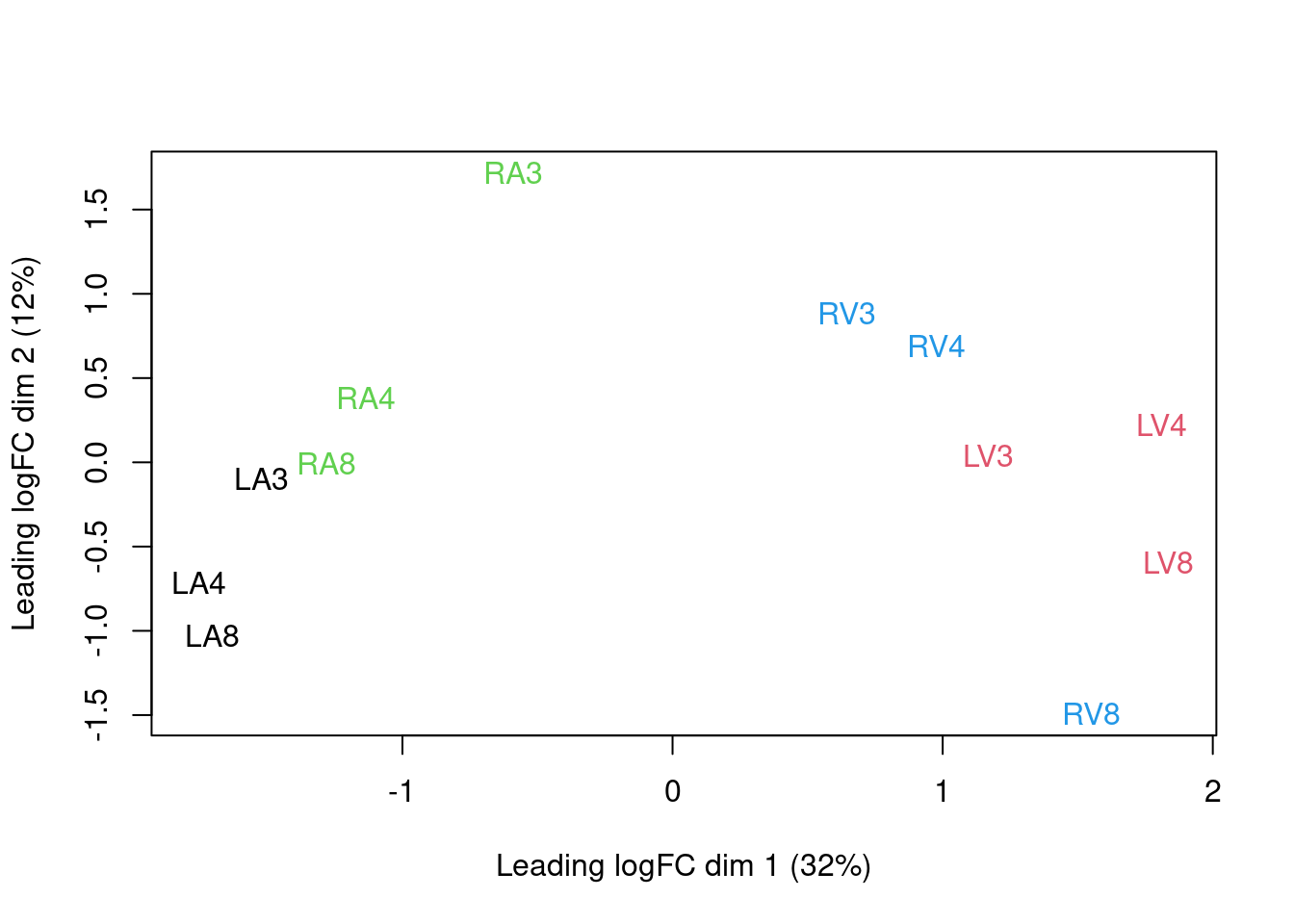
The first axis in the plot is showing the leading log fold changes (differences on the log scale) between the samples.
3.5 Summarization to protein level
We use robust summarization in aggregateFeatures. This is the default
workflow of aggregateFeatures so you do not have to specifiy the
argument fun. However, because we compare methods we have
included the fun argument to show the summarization method
explicitely.
pe <- aggregateFeatures(pe,
i = "peptideNorm",
fcol = "Proteins",
na.rm = TRUE,
name = "proteinRobust",
fun = MsCoreUtils::robustSummary)## Your quantitative and row data contain missing values. Please read the
## relevant section(s) in the aggregateFeatures manual page regarding the
## effects of missing values on data aggregation.plotMDS(assay(pe[["proteinRobust"]]),
col = colData(pe)$location:colData(pe)$tissue %>%
as.numeric,
labels = colData(pe) %>%
rownames %>%
substr(start = 11, stop = 13)
)
4 Data Analysis
4.1 Estimation
We model the protein level expression values using
msqrob. By default msqrob2 estimates the model
parameters using robust regression.
4.2 Inference
Explore Design
## $`location = L`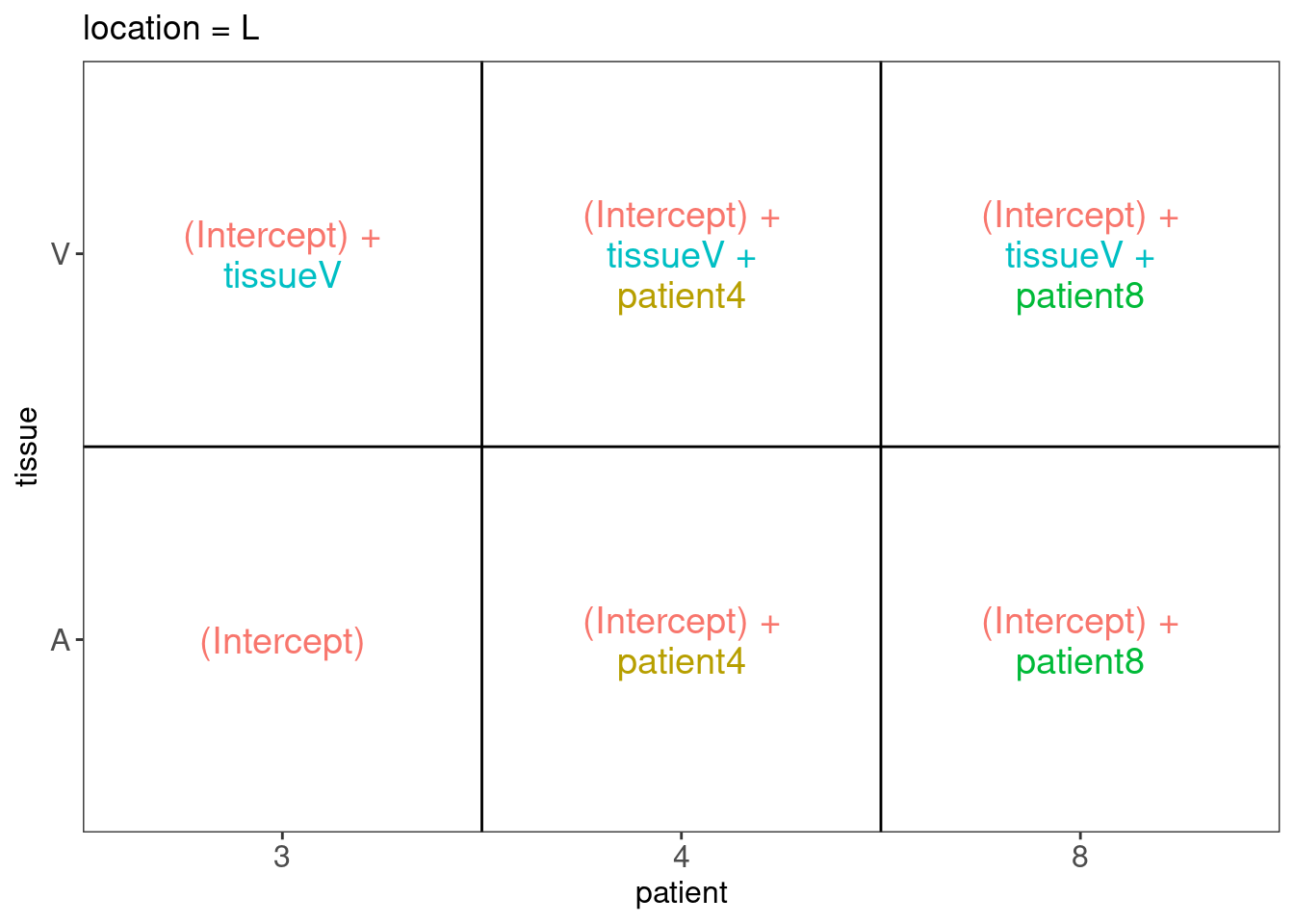
##
## $`location = R`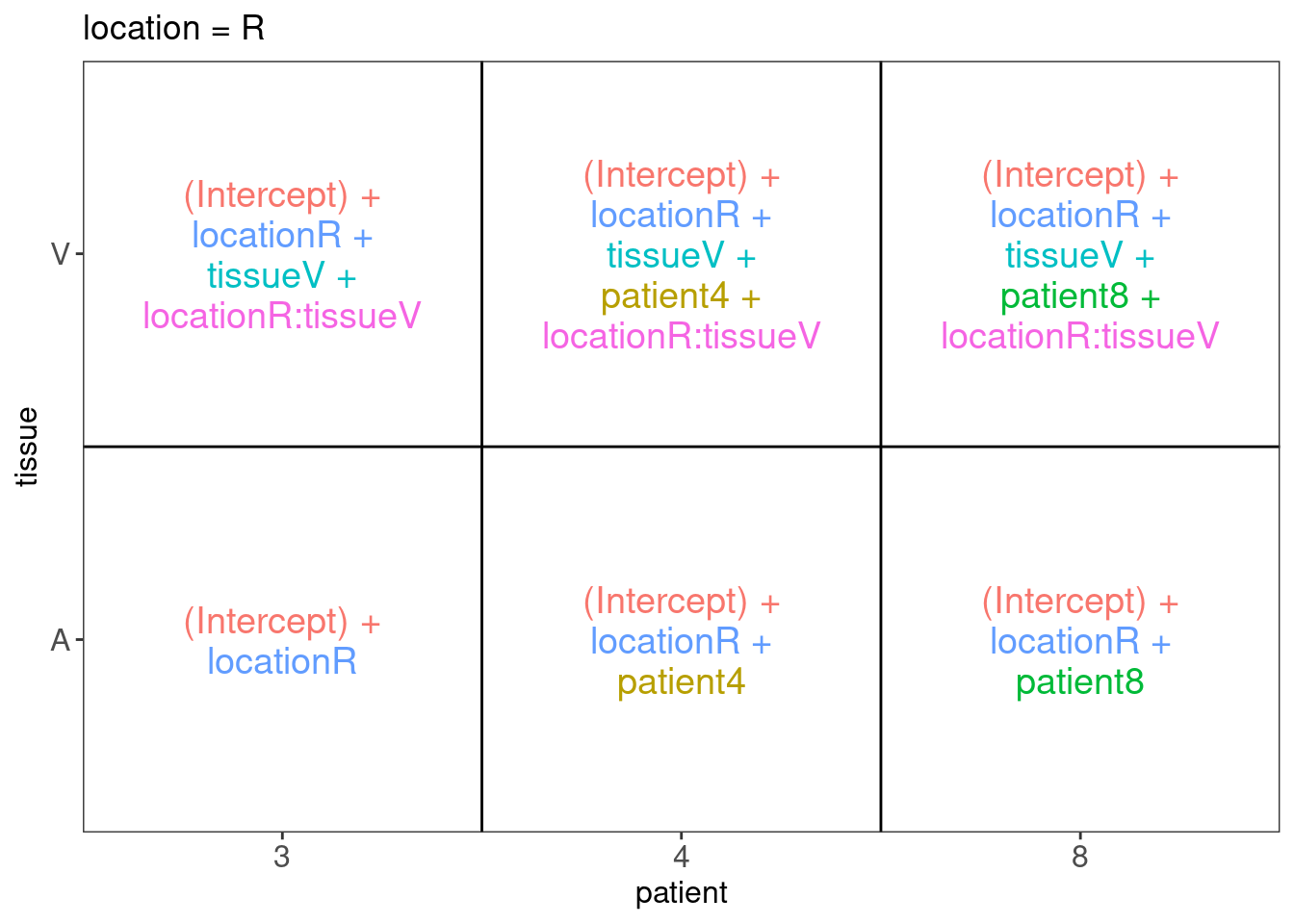
design <- model.matrix(~location*tissue + patient, data = colData(pe))
L <- makeContrast(
c(
"tissueV = 0",
"tissueV + locationR:tissueV = 0",
"tissueV + 0.5*locationR:tissueV = 0","locationR:tissueV = 0"),
parameterNames = colnames(design)
)
pe <- hypothesisTest(object = pe, i = "proteinRobust", contrast = L, overwrite=TRUE)4.3 Evaluate results contrast \(\log_2 FC_{V-A}^L\)
4.3.1 Volcano-plot
volcanoLeft <- ggplot(rowData(pe[["proteinRobust"]])$"tissueV",
aes(x = logFC, y = -log10(pval), color = adjPval < 0.05)) +
geom_point(cex = 2.5) +
scale_color_manual(values = alpha(c("black", "red"), 0.5)) + theme_minimal()
volcanoLeft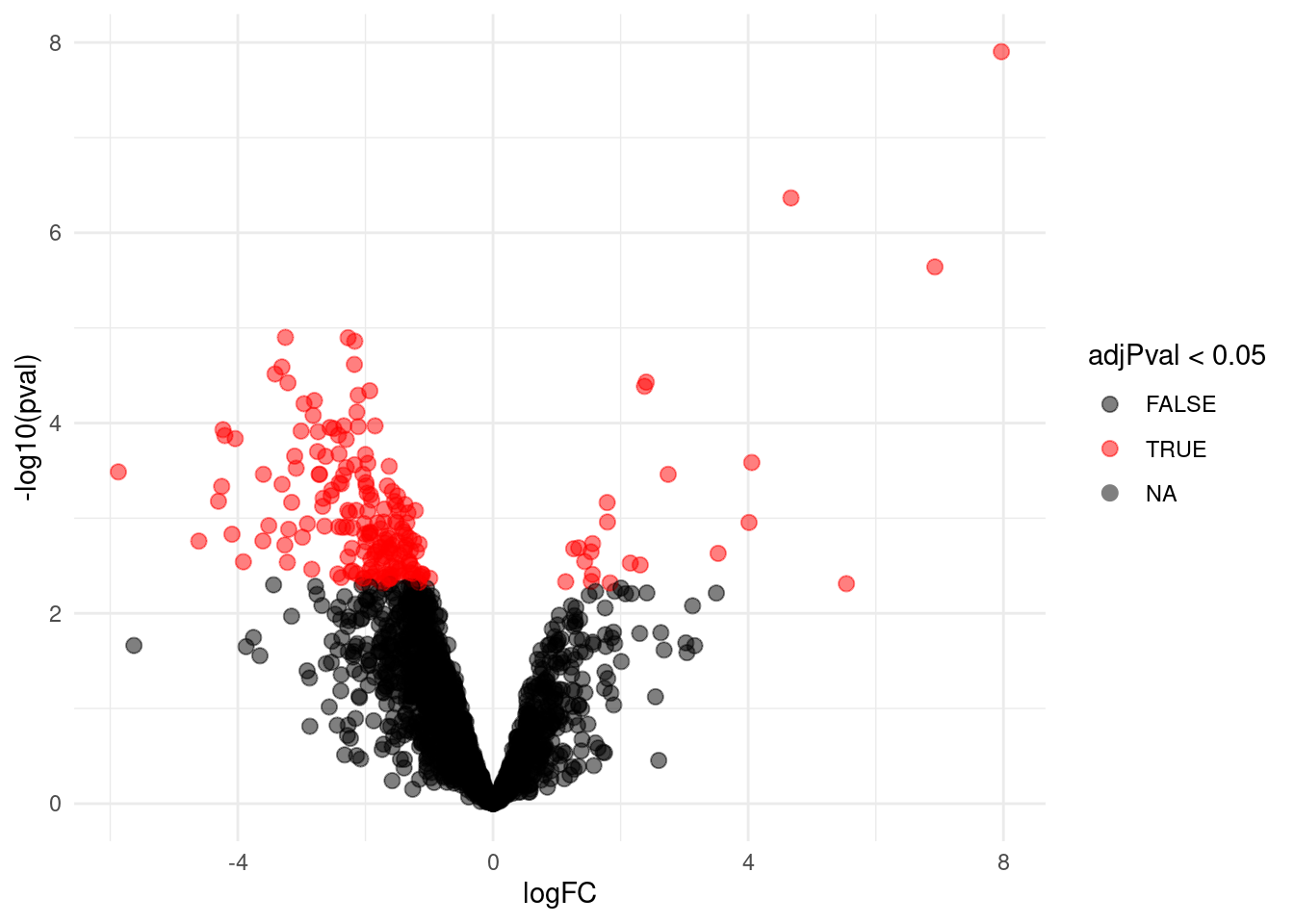
4.3.2 Heatmap
We first select the names of the proteins that were declared signficant.
sigNamesLeft <- rowData(pe[["proteinRobust"]])$tissueV %>%
rownames_to_column("proteinRobust") %>%
filter(adjPval<0.05) %>%
pull(proteinRobust)
heatmap(assay(pe[["proteinRobust"]])[sigNamesLeft, ])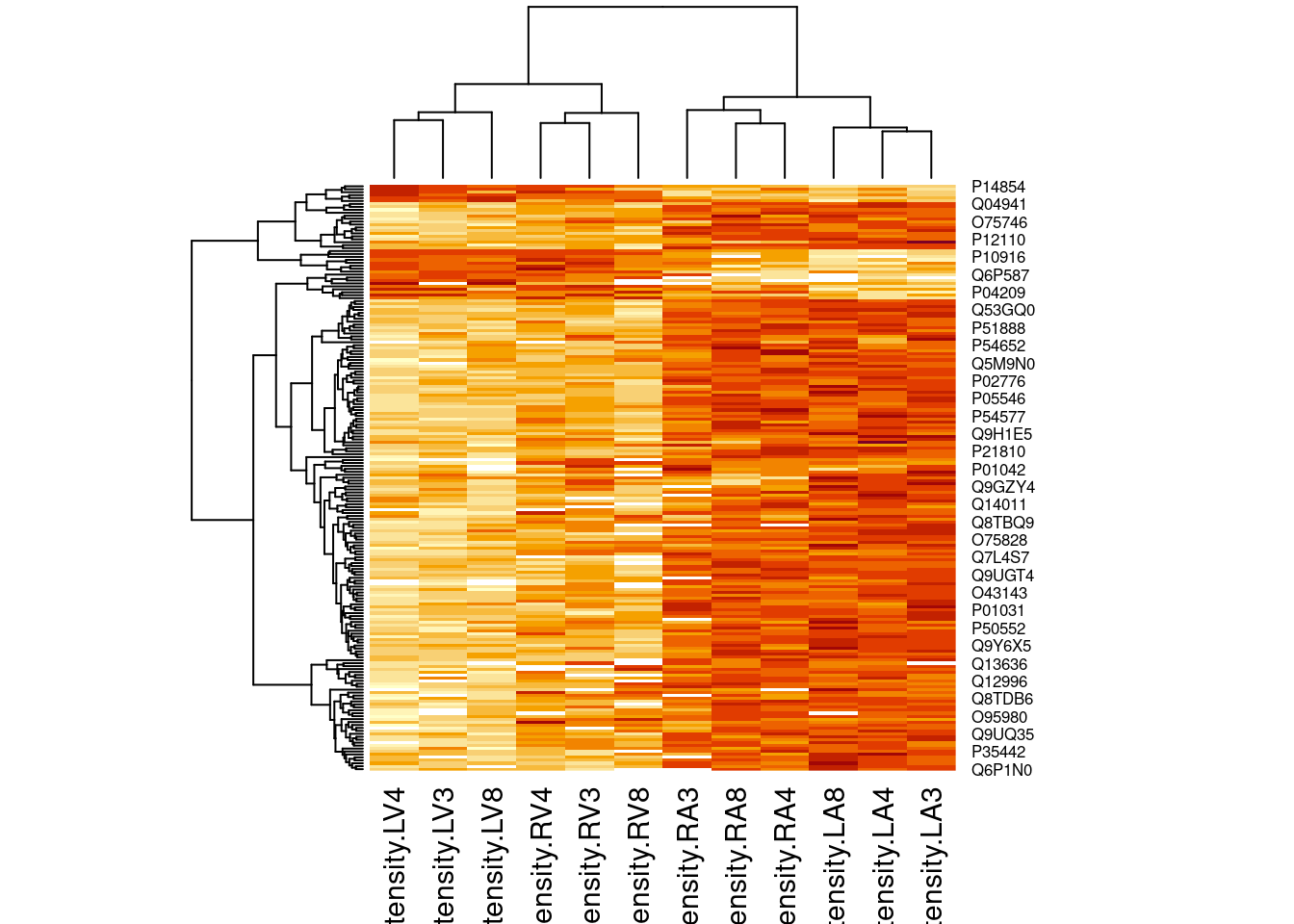
There are 205 proteins significantly differentially expressed at the 5% FDR level.
rowData(pe[["proteinRobust"]])$tissueV %>%
cbind(.,rowData(pe[["proteinRobust"]])$Protein.names) %>%
na.exclude %>%
filter(adjPval<0.05) %>%
arrange(pval) %>%
knitr::kable(.)| logFC | se | df | t | pval | adjPval | rowData(pe[[“proteinRobust”]])$Protein.names | |
|---|---|---|---|---|---|---|---|
| P08590 | 7.9679512 | 0.4386062 | 9.420359 | 18.166527 | 0.0000000 | 0.0000245 | Myosin light chain 3 |
| P12883 | 4.6698827 | 0.3735126 | 9.234880 | 12.502610 | 0.0000004 | 0.0004387 | Myosin-7 |
| P10916 | 6.9261087 | 0.5305670 | 7.420359 | 13.054165 | 0.0000022 | 0.0015039 | Myosin regulatory light chain 2, ventricular/cardiac muscle isoform |
| Q6UWY5 | -3.2536428 | 0.3846747 | 9.166472 | -8.458166 | 0.0000126 | 0.0050454 | Olfactomedin-like protein 1 |
| O75368 | -2.2715945 | 0.2746175 | 9.342472 | -8.271850 | 0.0000135 | 0.0050454 | SH3 domain-binding glutamic acid-rich-like protein |
| P46821 | -2.1668803 | 0.2664151 | 9.420359 | -8.133473 | 0.0000147 | 0.0050454 | Microtubule-associated protein 1B;MAP1B heavy chain;MAP1 light chain LC1 |
| O95865 | -2.1732192 | 0.2854548 | 9.420359 | -7.613183 | 0.0000254 | 0.0066381 | N(G),N(G)-dimethylarginine dimethylaminohydrolase 2 |
| Q8N474 | -3.3090235 | 0.3826720 | 7.950693 | -8.647153 | 0.0000258 | 0.0066381 | Secreted frizzled-related protein 1 |
| Q9ULL5-3 | -3.4165405 | 0.4016851 | 7.899339 | -8.505520 | 0.0000302 | 0.0069160 | Proline-rich protein 12 |
| P21810 | -3.2145420 | 0.4232686 | 8.810008 | -7.594568 | 0.0000376 | 0.0071804 | Biglycan |
| P14854 | 2.4019126 | 0.3085593 | 8.484681 | 7.784282 | 0.0000384 | 0.0071804 | Cytochrome c oxidase subunit 6B1 |
| O94875-10 | 2.3744000 | 0.3210591 | 8.958125 | 7.395523 | 0.0000423 | 0.0072492 | Sorbin and SH3 domain-containing protein 2 |
| P05546 | -1.9318824 | 0.2730471 | 9.324873 | -7.075271 | 0.0000486 | 0.0076991 | Heparin cofactor 2 |
| P29622 | -2.1134793 | 0.3039016 | 9.420359 | -6.954485 | 0.0000530 | 0.0077940 | Kallistatin |
| Q16647 | -2.7991521 | 0.3979468 | 9.081456 | -7.033986 | 0.0000582 | 0.0079891 | Prostacyclin synthase |
| P02452 | -2.9630153 | 0.4018280 | 8.345480 | -7.373839 | 0.0000628 | 0.0080762 | Collagen alpha-1(I) chain |
| P51884 | -2.1343314 | 0.3184899 | 9.210584 | -6.701409 | 0.0000793 | 0.0095563 | Lumican |
| Q8TBQ9 | -2.8189960 | 0.4079802 | 8.676104 | -6.909640 | 0.0000836 | 0.0095563 | Protein kish-A |
| P36955 | -2.3377901 | 0.3380798 | 8.196767 | -6.914907 | 0.0001093 | 0.0097605 | Pigment epithelium-derived factor |
| Q9UBG0 | -2.5547510 | 0.3990636 | 9.225608 | -6.401864 | 0.0001120 | 0.0097605 | C-type mannose receptor 2 |
| P00325 | -2.1119462 | 0.3259838 | 9.037213 | -6.478684 | 0.0001122 | 0.0097605 | Alcohol dehydrogenase 1B |
| P07451 | -1.8489373 | 0.2923533 | 9.420359 | -6.324324 | 0.0001123 | 0.0097605 | Carbonic anhydrase 3 |
| P23083 | -4.2325442 | 0.5743071 | 7.420359 | -7.369827 | 0.0001148 | 0.0097605 | Ig heavy chain V-I region V35 |
| P24844 | -2.4962161 | 0.3945586 | 9.365146 | -6.326604 | 0.0001149 | 0.0097605 | Myosin regulatory light polypeptide 9 |
| Q15113 | -2.7424988 | 0.4339863 | 9.239033 | -6.319320 | 0.0001230 | 0.0097605 | Procollagen C-endopeptidase enhancer 1 |
| P51888 | -3.0081810 | 0.4037624 | 7.212550 | -7.450374 | 0.0001233 | 0.0097605 | Prolargin |
| Q06828 | -4.2030186 | 0.6620572 | 9.022216 | -6.348422 | 0.0001317 | 0.0099192 | Fibromodulin |
| Q53GQ0 | -2.4232423 | 0.3921550 | 9.410782 | -6.179296 | 0.0001350 | 0.0099192 | Very-long-chain 3-oxoacyl-CoA reductase |
| P13533 | -4.0404076 | 0.6480615 | 9.151585 | -6.234605 | 0.0001420 | 0.0100740 | Myosin-6 |
| P35442 | -2.3010500 | 0.3542666 | 8.420359 | -6.495250 | 0.0001509 | 0.0103503 | Thrombospondin-2 |
| P08294 | -2.7503241 | 0.4275819 | 8.014829 | -6.432274 | 0.0002005 | 0.0131731 | Extracellular superoxide dismutase [Cu-Zn] |
| Q96LL9 | -2.4127080 | 0.4001370 | 8.810329 | -6.029705 | 0.0002128 | 0.0131731 | DnaJ homolog subfamily C member 30 |
| P36021 | -3.1085951 | 0.5045027 | 8.420359 | -6.161702 | 0.0002193 | 0.0131731 | Monocarboxylate transporter 8 |
| P18428 | -1.9996058 | 0.3407167 | 9.191871 | -5.868822 | 0.0002193 | 0.0131731 | Lipopolysaccharide-binding protein |
| Q9UL18 | -2.6225313 | 0.4111229 | 7.913528 | -6.378947 | 0.0002240 | 0.0131731 | Protein argonaute-1 |
| Q92508 | 4.0552373 | 0.5586150 | 6.420359 | 7.259450 | 0.0002541 | 0.0145255 | Piezo-type mechanosensitive ion channel component 1 |
| P46060 | -1.9620904 | 0.3284336 | 8.420359 | -5.974085 | 0.0002723 | 0.0150027 | Ran GTPase-activating protein 1 |
| P02743 | -2.1700557 | 0.3863985 | 9.420359 | -5.616108 | 0.0002772 | 0.0150027 | Serum amyloid P-component;Serum amyloid P-component(1-203) |
| P05997 | -3.0856148 | 0.5411521 | 8.993385 | -5.701936 | 0.0002944 | 0.0150027 | Collagen alpha-2(V) chain |
| Q9UGT4 | -2.3006162 | 0.4092617 | 9.245036 | -5.621381 | 0.0002949 | 0.0150027 | Sushi domain-containing protein 2 |
| Q14764 | -1.6267934 | 0.2926175 | 9.420359 | -5.559454 | 0.0002989 | 0.0150027 | Major vault protein |
| Q8WWA0 | -5.8725166 | 0.9857040 | 8.156016 | -5.957687 | 0.0003145 | 0.0151581 | Intelectin-1 |
| O60760 | -3.5990151 | 0.6219772 | 8.420359 | -5.786410 | 0.0003396 | 0.0151581 | Hematopoietic prostaglandin D synthase |
| O43677 | -2.7199107 | 0.4724234 | 8.483808 | -5.757358 | 0.0003418 | 0.0151581 | NADH dehydrogenase [ubiquinone] 1 subunit C1, mitochondrial |
| Q9BW30 | -2.7245919 | 0.4899368 | 9.052429 | -5.561109 | 0.0003441 | 0.0151581 | Tubulin polymerization-promoting protein family member 3 |
| Q9P2B2 | -2.0393915 | 0.3747097 | 9.420359 | -5.442591 | 0.0003496 | 0.0151581 | Prostaglandin F2 receptor negative regulator |
| Q9UBB5 | 2.7451793 | 0.4000606 | 6.420359 | 6.861908 | 0.0003517 | 0.0151581 | Methyl-CpG-binding domain protein 2 |
| P40261 | -2.3445730 | 0.4282291 | 9.273912 | -5.475044 | 0.0003535 | 0.0151581 | Nicotinamide N-methyltransferase |
| Q9NZ01 | -2.4109216 | 0.4375281 | 8.685989 | -5.510325 | 0.0004252 | 0.0172070 | Very-long-chain enoyl-CoA reductase |
| P00748 | -1.9914680 | 0.3712703 | 9.156315 | -5.363931 | 0.0004282 | 0.0172070 | Coagulation factor XII;Coagulation factor XIIa heavy chain;Beta-factor XIIa part 1;Coagulation factor XIIa light chain |
| Q92736-2 | -3.3064395 | 0.6042970 | 8.766703 | -5.471547 | 0.0004323 | 0.0172070 | Ryanodine receptor 2 |
| O14967 | -2.3785961 | 0.4303723 | 8.580509 | -5.526834 | 0.0004348 | 0.0172070 | Calmegin |
| O00180 | -4.2526703 | 0.7658995 | 8.420359 | -5.552517 | 0.0004502 | 0.0174825 | Potassium channel subfamily K member 1 |
| P12110 | -1.9881397 | 0.3554165 | 8.222232 | -5.593831 | 0.0004662 | 0.0177279 | Collagen alpha-2(VI) chain |
| Q07954 | -1.6573532 | 0.3175193 | 9.420359 | -5.219693 | 0.0004738 | 0.0177279 | Prolow-density lipoprotein receptor-related protein 1;Low-density lipoprotein receptor-related protein 1 85 kDa subunit;Low-density lipoprotein receptor-related protein 1 515 kDa subunit;Low-density lipoprotein receptor-related protein 1 intracellular domain |
| O95980 | -2.5277905 | 0.4569818 | 8.199718 | -5.531490 | 0.0005071 | 0.0186373 | Reversion-inducing cysteine-rich protein with Kazal motifs |
| P31994-2 | -1.5795166 | 0.3043264 | 9.122267 | -5.190206 | 0.0005473 | 0.0196543 | Low affinity immunoglobulin gamma Fc region receptor II-b |
| O00264 | -1.9731043 | 0.3803391 | 9.097380 | -5.187750 | 0.0005539 | 0.0196543 | Membrane-associated progesterone receptor component 1 |
| Q14195-2 | -2.5380012 | 0.4966607 | 9.283388 | -5.110131 | 0.0005775 | 0.0198432 | Dihydropyrimidinase-related protein 3 |
| Q8TBP6 | -1.9233155 | 0.3718274 | 9.032046 | -5.172602 | 0.0005785 | 0.0198432 | Solute carrier family 25 member 40 |
| Q8WZA9 | -1.4960398 | 0.2943543 | 9.231251 | -5.082446 | 0.0006106 | 0.0203049 | Immunity-related GTPase family Q protein |
| Q9BXN1 | -2.6634064 | 0.5185471 | 9.012323 | -5.136287 | 0.0006117 | 0.0203049 | Asporin |
| O43464 | -1.9028432 | 0.3769428 | 9.166698 | -5.048095 | 0.0006542 | 0.0210534 | Serine protease HTRA2, mitochondrial |
| P01699 | -4.3035389 | 0.6908090 | 6.301890 | -6.229709 | 0.0006547 | 0.0210534 | Ig lambda chain V-I region VOR |
| Q9GZY4 | -3.1557642 | 0.5788229 | 7.775949 | -5.452037 | 0.0006690 | 0.0211457 | Cytochrome c oxidase assembly factor 1 homolog |
| Q9NVN8 | -6.0616246 | 0.8752359 | 5.420359 | -6.925704 | 0.0006911 | 0.0211457 | Guanine nucleotide-binding protein-like 3-like protein |
| P41240 | -1.5466953 | 0.3073684 | 9.050310 | -5.032058 | 0.0006955 | 0.0211457 | Tyrosine-protein kinase CSK |
| P06858 | 1.7891922 | 0.3600895 | 9.302479 | 4.968743 | 0.0006987 | 0.0211457 | Lipoprotein lipase |
| O95631 | -3.9795086 | 0.5835987 | 5.420359 | -6.818912 | 0.0007462 | 0.0217213 | Netrin-1 |
| P04083 | -1.5201990 | 0.3111586 | 9.420359 | -4.885608 | 0.0007573 | 0.0217213 | Annexin A1 |
| Q8NAT1 | -1.3792038 | 0.2816741 | 9.360836 | -4.896452 | 0.0007596 | 0.0217213 | Protein O-linked-mannose beta-1,4-N-acetylglucosaminyltransferase 2 |
| P02747 | -2.6697776 | 0.4727594 | 7.043683 | -5.647223 | 0.0007599 | 0.0217213 | Complement C1q subcomponent subunit C |
| Q92621 | -1.7036447 | 0.3526893 | 9.417394 | -4.830440 | 0.0008203 | 0.0227488 | Nuclear pore complex protein Nup205 |
| A6NMZ7 | -2.2762344 | 0.4717832 | 9.420359 | -4.824746 | 0.0008263 | 0.0227488 | Collagen alpha-6(VI) chain |
| Q9NY15 | -2.1436460 | 0.4441922 | 9.403688 | -4.825943 | 0.0008290 | 0.0227488 | Stabilin-1 |
| P02775 | -1.9622566 | 0.4084962 | 9.420359 | -4.803610 | 0.0008518 | 0.0230658 | Platelet basic protein;Connective tissue-activating peptide III;TC-2;Connective tissue-activating peptide III(1-81);Beta-thromboglobulin;Neutrophil-activating peptide 2(74);Neutrophil-activating peptide 2(73);Neutrophil-activating peptide 2;TC-1;Neutrophil-activating peptide 2(1-66);Neutrophil-activating peptide 2(1-63) |
| Q92604 | -2.2504519 | 0.4687530 | 9.352385 | -4.800934 | 0.0008727 | 0.0233221 | Acyl-CoA:lysophosphatidylglycerol acyltransferase 1 |
| Q96C86 | -1.4762676 | 0.3054387 | 9.149198 | -4.833270 | 0.0008871 | 0.0233221 | m7GpppX diphosphatase |
| O14980 | -1.2169774 | 0.2551792 | 9.420359 | -4.769109 | 0.0008953 | 0.0233221 | Exportin-1 |
| P48681 | -1.3334013 | 0.2810861 | 9.417394 | -4.743747 | 0.0009295 | 0.0239120 | Nestin |
| Q9UNW9 | 4.0122337 | 0.8547106 | 9.142176 | 4.694260 | 0.0010823 | 0.0268544 | RNA-binding protein Nova-2 |
| P24311 | 1.7937075 | 0.3882030 | 9.420359 | 4.620540 | 0.0011114 | 0.0268544 | Cytochrome c oxidase subunit 7B, mitochondrial |
| P50552 | -1.5288544 | 0.3255300 | 9.036834 | -4.696508 | 0.0011133 | 0.0268544 | Vasodilator-stimulated phosphoprotein |
| Q53GG5-2 | -2.9088697 | 0.6018399 | 8.409844 | -4.833295 | 0.0011275 | 0.0268544 | PDZ and LIM domain protein 3 |
| Q7L4S7 | -1.7124303 | 0.3273125 | 7.153052 | -5.231790 | 0.0011302 | 0.0268544 | Protein ARMCX6 |
| Q8WY22 | -1.8091979 | 0.3749461 | 8.420359 | -4.825221 | 0.0011356 | 0.0268544 | BRI3-binding protein |
| P56539 | -2.0161968 | 0.4222161 | 8.599057 | -4.775273 | 0.0011445 | 0.0268544 | Caveolin-3 |
| P49207 | -1.5105253 | 0.3230060 | 9.024427 | -4.676462 | 0.0011496 | 0.0268544 | 60S ribosomal protein L34 |
| Q5M9N0 | -3.5146838 | 0.7308851 | 8.385642 | -4.808805 | 0.0011741 | 0.0268544 | Coiled-coil domain-containing protein 158 |
| P46063 | -1.3496984 | 0.2916572 | 9.183164 | -4.627688 | 0.0011764 | 0.0268544 | ATP-dependent DNA helicase Q1 |
| Q6SZW1 | -2.6395602 | 0.5203026 | 7.420359 | -5.073125 | 0.0012090 | 0.0268544 | Sterile alpha and TIR motif-containing protein 1 |
| P02671 | -2.1346012 | 0.3881675 | 6.417394 | -5.499175 | 0.0012144 | 0.0268544 | Fibrinogen alpha chain;Fibrinopeptide A;Fibrinogen alpha chain |
| Q5NDL2 | -2.2948242 | 0.4886297 | 8.717721 | -4.696449 | 0.0012273 | 0.0268544 | EGF domain-specific O-linked N-acetylglucosamine transferase |
| Q9HAV4 | -2.4188578 | 0.4860155 | 7.670496 | -4.976915 | 0.0012275 | 0.0268544 | Exportin-5 |
| Q9BXR6 | -2.3571850 | 0.5184175 | 9.417394 | -4.546886 | 0.0012396 | 0.0268544 | Complement factor H-related protein 5 |
| Q92681 | -2.1962163 | 0.4515818 | 7.932336 | -4.863385 | 0.0012812 | 0.0274650 | Regulatory solute carrier protein family 1 member 1 |
| Q96H79 | -3.2031583 | 0.5775193 | 6.200924 | -5.546409 | 0.0013005 | 0.0275566 | Zinc finger CCCH-type antiviral protein 1-like |
| Q8TDB6 | -1.7610737 | 0.3855338 | 9.102046 | -4.567884 | 0.0013122 | 0.0275566 | E3 ubiquitin-protein ligase DTX3L |
| O15230 | -1.4246624 | 0.3071116 | 8.689548 | -4.638909 | 0.0013403 | 0.0278624 | Laminin subunit alpha-5 |
| Q15274 | -1.9285076 | 0.4023627 | 7.916098 | -4.792959 | 0.0014087 | 0.0289903 | Nicotinate-nucleotide pyrophosphorylase [carboxylating] |
| Q9BUF5 | -1.9157084 | 0.4285093 | 9.301065 | -4.470635 | 0.0014314 | 0.0290457 | Tubulin beta-6 chain |
| Q9Y5U8 | -4.0913780 | 0.9166412 | 9.319580 | -4.463445 | 0.0014396 | 0.0290457 | Mitochondrial pyruvate carrier 1 |
| P14550 | -1.3982496 | 0.3076929 | 8.855961 | -4.544302 | 0.0014562 | 0.0290948 | Alcohol dehydrogenase [NADP(+)] |
| O15111 | -1.9484725 | 0.3851415 | 6.956358 | -5.059108 | 0.0014925 | 0.0295349 | Inhibitor of nuclear factor kappa-B kinase subunit alpha |
| Q12996 | -2.0094808 | 0.3831587 | 6.493841 | -5.244512 | 0.0015107 | 0.0296094 | Cleavage stimulation factor subunit 3 |
| Q9UBI9 | -2.9866384 | 0.6067195 | 7.231219 | -4.922602 | 0.0015558 | 0.0296875 | Headcase protein homolog |
| O75828 | -1.6544202 | 0.3767450 | 9.420359 | -4.391353 | 0.0015612 | 0.0296875 | Carbonyl reductase [NADPH] 3 |
| P06727 | -1.3683135 | 0.3043718 | 8.840955 | -4.495533 | 0.0015671 | 0.0296875 | Apolipoprotein A-IV |
| P04196 | -1.9303128 | 0.4001227 | 7.506343 | -4.824303 | 0.0015724 | 0.0296875 | Histidine-rich glycoprotein |
| Q9ULC3 | -1.6333020 | 0.3645496 | 8.720682 | -4.480329 | 0.0016570 | 0.0309524 | Ras-related protein Rab-23 |
| P14555 | -4.6109150 | 0.9289708 | 6.916563 | -4.963466 | 0.0016891 | 0.0309524 | Phospholipase A2, membrane associated |
| P02461 | -3.6062939 | 0.7964105 | 8.416773 | -4.528185 | 0.0016954 | 0.0309524 | Collagen alpha-1(III) chain |
| Q6ZSY5 | -1.6890696 | 0.3871892 | 9.235896 | -4.362389 | 0.0017089 | 0.0309524 | Protein phosphatase 1 regulatory subunit 3F |
| P01031 | -1.3157564 | 0.2928900 | 8.544930 | -4.492322 | 0.0017146 | 0.0309524 | Complement C5;Complement C5 beta chain;Complement C5 alpha chain;C5a anaphylatoxin;Complement C5 alpha chain |
| Q8N5M1 | -1.9899041 | 0.4567857 | 9.122947 | -4.356319 | 0.0017753 | 0.0317701 | ATP synthase mitochondrial F1 complex assembly factor 2 |
| P04003 | -1.5009188 | 0.3436224 | 9.023047 | -4.367931 | 0.0017916 | 0.0317849 | C4b-binding protein alpha chain |
| P45877 | -1.2442932 | 0.2899715 | 9.420359 | -4.291088 | 0.0018158 | 0.0319386 | Peptidyl-prolyl cis-trans isomerase C |
| P08582 | -1.6911943 | 0.3948150 | 9.394818 | -4.283511 | 0.0018482 | 0.0322331 | Melanotransferrin |
| Q6YN16 | 1.5603531 | 0.3657250 | 9.420359 | 4.266465 | 0.0018848 | 0.0325586 | Hydroxysteroid dehydrogenase-like protein 2 |
| Q13636 | -3.2633858 | 0.6344956 | 6.233646 | -5.143276 | 0.0018985 | 0.0325586 | Ras-related protein Rab-31 |
| O75746 | -1.5795706 | 0.3608149 | 8.652741 | -4.377787 | 0.0019522 | 0.0332039 | Calcium-binding mitochondrial carrier protein Aralar1 |
| Q9UQ35 | -1.3570787 | 0.3202712 | 9.401099 | -4.237279 | 0.0019793 | 0.0332620 | Serine/arginine repetitive matrix protein 2 |
| P34932 | -1.1578310 | 0.2723151 | 9.291753 | -4.251806 | 0.0019880 | 0.0332620 | Heat shock 70 kDa protein 4 |
| Q5VIR6-4 | -1.7638024 | 0.4142915 | 9.167220 | -4.257395 | 0.0020325 | 0.0336262 | Vacuolar protein sorting-associated protein 53 homolog |
| P04004 | -1.6269790 | 0.3774162 | 8.785099 | -4.310835 | 0.0020725 | 0.0336262 | Vitronectin;Vitronectin V65 subunit;Vitronectin V10 subunit;Somatomedin-B |
| P01042 | -2.2103015 | 0.4751169 | 7.306380 | -4.652122 | 0.0020862 | 0.0336262 | Kininogen-1;Kininogen-1 heavy chain;T-kinin;Bradykinin;Lysyl-bradykinin;Kininogen-1 light chain;Low molecular weight growth-promoting factor |
| P02776 | -1.7615098 | 0.3949258 | 8.029185 | -4.460356 | 0.0020911 | 0.0336262 | Platelet factor 4;Platelet factor 4, short form |
| Q9H1E5 | -1.3852786 | 0.3303392 | 9.420359 | -4.193504 | 0.0021061 | 0.0336262 | Thioredoxin-related transmembrane protein 4 |
| Q8WWQ0 | -1.6224614 | 0.3871144 | 9.420359 | -4.191168 | 0.0021137 | 0.0336262 | PH-interacting protein |
| P04209 | 1.3436120 | 0.3180816 | 9.190380 | 4.224111 | 0.0021241 | 0.0336262 | Ig lambda chain V-II region NIG-84 |
| P15924 | 1.2648885 | 0.3029535 | 9.383746 | 4.175190 | 0.0021844 | 0.0343167 | Desmoplakin |
| Q9BTV4 | -1.8099873 | 0.4323541 | 9.211817 | -4.186353 | 0.0022365 | 0.0344787 | Transmembrane protein 43 |
| Q08945 | -2.0204841 | 0.4675424 | 8.420359 | -4.321499 | 0.0022538 | 0.0344787 | FACT complex subunit SSRP1 |
| Q2TAA5 | -1.2916274 | 0.3093292 | 9.234456 | -4.175575 | 0.0022609 | 0.0344787 | GDP-Man:Man(3)GlcNAc(2)-PP-Dol alpha-1,2-mannosyltransferase |
| Q53T59 | -1.8461756 | 0.4200861 | 8.037638 | -4.394755 | 0.0022769 | 0.0344787 | HCLS1-binding protein 3 |
| Q04721 | 1.5358466 | 0.3685976 | 9.213519 | 4.166730 | 0.0023027 | 0.0344787 | Neurogenic locus notch homolog protein 2;Notch 2 extracellular truncation;Notch 2 intracellular domain |
| P09619 | -1.6085799 | 0.3866754 | 9.226613 | -4.160027 | 0.0023189 | 0.0344787 | Platelet-derived growth factor receptor beta |
| Q5JPH6 | 3.5281305 | 0.6863030 | 5.829289 | 5.140777 | 0.0023262 | 0.0344787 | Probable glutamate–tRNA ligase, mitochondrial |
| P05455 | -1.2004499 | 0.2883779 | 9.171164 | -4.162767 | 0.0023398 | 0.0344787 | Lupus La protein |
| Q9Y6X5 | -1.3875045 | 0.3272759 | 8.705094 | -4.239556 | 0.0023455 | 0.0344787 | Bis(5-adenosyl)-triphosphatase ENPP4 |
| O60262 | -1.5577722 | 0.3689381 | 8.700017 | -4.222313 | 0.0024077 | 0.0351428 | Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-7 |
| O75348 | -1.8591253 | 0.4535299 | 9.420359 | -4.099234 | 0.0024337 | 0.0352720 | V-type proton ATPase subunit G 1 |
| O00303 | -1.4561773 | 0.3518679 | 9.051187 | -4.138420 | 0.0024969 | 0.0359346 | Eukaryotic translation initiation factor 3 subunit F |
| P01034 | -1.6978634 | 0.4087932 | 8.859133 | -4.153355 | 0.0025581 | 0.0365590 | Cystatin-C |
| P49458 | -2.2735269 | 0.5075335 | 7.251835 | -4.479560 | 0.0026268 | 0.0372073 | Signal recognition particle 9 kDa protein |
| P25940 | -1.7454125 | 0.4272441 | 9.155619 | -4.085281 | 0.0026396 | 0.0372073 | Collagen alpha-3(V) chain |
| P62760 | -1.9146688 | 0.4707491 | 9.159766 | -4.067281 | 0.0027100 | 0.0379401 | Visinin-like protein 1 |
| Q14019 | -3.9129083 | 0.8333648 | 6.420359 | -4.695313 | 0.0027954 | 0.0388713 | Coactosin-like protein |
| O15116 | -3.2231772 | 0.6507124 | 5.794630 | -4.953305 | 0.0028355 | 0.0391643 | U6 snRNA-associated Sm-like protein LSm1 |
| Q9UKS6 | 1.4372286 | 0.3556788 | 8.991910 | 4.040805 | 0.0029305 | 0.0398449 | Protein kinase C and casein kinase substrate in neurons protein 3 |
| P08311 | -1.3139510 | 0.3238932 | 8.882238 | -4.056741 | 0.0029350 | 0.0398449 | Cathepsin G |
| Q9UKX3 | 2.1517579 | 0.5377017 | 9.238422 | 4.001769 | 0.0029429 | 0.0398449 | Myosin-13 |
| P30405 | -1.5177122 | 0.3822973 | 9.420359 | -3.969979 | 0.0029732 | 0.0399918 | Peptidyl-prolyl cis-trans isomerase F, mitochondrial |
| P36551 | -1.3202398 | 0.3291895 | 9.078769 | -4.010577 | 0.0030076 | 0.0401444 | Oxygen-dependent coproporphyrinogen-III oxidase, mitochondrial |
| Q9Y2Z0 | -1.7414076 | 0.4398381 | 9.420359 | -3.959201 | 0.0030235 | 0.0401444 | Suppressor of G2 allele of SKP1 homolog |
| P19429 | 2.3063639 | 0.5572199 | 8.211191 | 4.139056 | 0.0030787 | 0.0406150 | Troponin I, cardiac muscle |
| P54577 | -1.3002728 | 0.3261820 | 9.016679 | -3.986341 | 0.0031635 | 0.0414677 | Tyrosine–tRNA ligase, cytoplasmic;Tyrosine–tRNA ligase, cytoplasmic, N-terminally processed |
| P54652 | -1.5932853 | 0.4025031 | 9.131756 | -3.958443 | 0.0032184 | 0.0419203 | Heat shock-related 70 kDa protein 2 |
| O75475 | -1.9038879 | 0.4846934 | 9.293000 | -3.928025 | 0.0032591 | 0.0421839 | PC4 and SFRS1-interacting protein |
| Q7LBR1 | -2.8413335 | 0.7311661 | 9.420359 | -3.886030 | 0.0033900 | 0.0434941 | Charged multivesicular body protein 1b |
| P25311 | -1.5051007 | 0.3754727 | 8.556242 | -4.008549 | 0.0034026 | 0.0434941 | Zinc-alpha-2-glycoprotein |
| P60468 | -1.4647431 | 0.3625247 | 8.275558 | -4.040396 | 0.0034805 | 0.0442150 | Protein transport protein Sec61 subunit beta |
| Q04941 | -1.9356392 | 0.4958892 | 9.122186 | -3.903371 | 0.0035088 | 0.0443012 | Proteolipid protein 2 |
| Q9BS26 | -1.2463986 | 0.3225161 | 9.354832 | -3.864609 | 0.0035522 | 0.0443625 | Endoplasmic reticulum resident protein 44 |
| P03950 | -2.2030636 | 0.5456390 | 8.205145 | -4.037585 | 0.0035568 | 0.0443625 | Angiogenin |
| Q13641 | -2.2297004 | 0.5236478 | 7.111870 | -4.258015 | 0.0036223 | 0.0449080 | Trophoblast glycoprotein |
| Q1KMD3 | -1.4744120 | 0.3796403 | 9.065422 | -3.883707 | 0.0036596 | 0.0450983 | Heterogeneous nuclear ribonucleoprotein U-like protein 2 |
| P26447 | -1.8372154 | 0.4683842 | 8.758554 | -3.922454 | 0.0036879 | 0.0451764 | Protein S100-A4 |
| Q96PK6 | -1.3089284 | 0.3399205 | 9.118686 | -3.850690 | 0.0038077 | 0.0461431 | RNA-binding protein 14 |
| O00567 | -2.4335450 | 0.6369246 | 9.345669 | -3.820774 | 0.0038116 | 0.0461431 | Nucleolar protein 56 |
| P12814 | -1.6827515 | 0.4236455 | 8.209855 | -3.972075 | 0.0038987 | 0.0465538 | Alpha-actinin-1 |
| Q9Y3B4 | -1.2603707 | 0.3283823 | 9.068421 | -3.838121 | 0.0039223 | 0.0465538 | Splicing factor 3B subunit 6 |
| Q86VU5 | 1.5583619 | 0.4110443 | 9.420359 | 3.791226 | 0.0039358 | 0.0465538 | Catechol O-methyltransferase domain-containing protein 1 |
| Q13478 | -1.6083281 | 0.4182581 | 8.998109 | -3.845301 | 0.0039360 | 0.0465538 | Interleukin-18 receptor 1 |
| Q9HB40 | -2.1393191 | 0.4860845 | 6.361260 | -4.401126 | 0.0039678 | 0.0466612 | Retinoid-inducible serine carboxypeptidase |
| P01024 | -1.1180307 | 0.2936124 | 9.157785 | -3.807846 | 0.0040365 | 0.0469892 | Complement C3;Complement C3 beta chain;C3-beta-c;Complement C3 alpha chain;C3a anaphylatoxin;Acylation stimulating protein;Complement C3b alpha chain;Complement C3c alpha chain fragment 1;Complement C3dg fragment;Complement C3g fragment;Complement C3d fragment;Complement C3f fragment;Complement C3c alpha chain fragment 2 |
| P01008 | -1.5692162 | 0.4151767 | 9.376569 | -3.779635 | 0.0040423 | 0.0469892 | Antithrombin-III |
| P07384 | -1.1052065 | 0.2922209 | 9.301416 | -3.782093 | 0.0040852 | 0.0469892 | Calpain-1 catalytic subunit |
| P14543 | -1.4365423 | 0.3779966 | 9.153301 | -3.800411 | 0.0040870 | 0.0469892 | Nidogen-1 |
| Q14011 | -2.3803571 | 0.6080384 | 8.287145 | -3.914814 | 0.0041543 | 0.0469943 | Cold-inducible RNA-binding protein |
| Q9UK22 | -1.8807802 | 0.4332817 | 6.412167 | -4.340779 | 0.0041699 | 0.0469943 | F-box only protein 2 |
| Q7Z3T8 | -1.6269924 | 0.4212289 | 8.586144 | -3.862490 | 0.0041889 | 0.0469943 | Zinc finger FYVE domain-containing protein 16 |
| O95486 | -1.7001274 | 0.4483551 | 9.078202 | -3.791922 | 0.0042035 | 0.0469943 | Protein transport protein Sec24A |
| O94919 | -1.1247209 | 0.2979363 | 9.184104 | -3.775038 | 0.0042258 | 0.0469943 | Endonuclease domain-containing 1 protein |
| Q9Y2D4 | -2.0300937 | 0.5292407 | 8.722544 | -3.835861 | 0.0042314 | 0.0469943 | Exocyst complex component 6B |
| Q9BXY0 | -1.9170100 | 0.4391665 | 6.280367 | -4.365109 | 0.0042597 | 0.0469943 | Protein MAK16 homolog |
| P07357 | -1.1425305 | 0.3041753 | 9.262894 | -3.756158 | 0.0042866 | 0.0469943 | Complement component C8 alpha chain |
| Q96ST3 | -1.6077298 | 0.4259265 | 9.106665 | -3.774665 | 0.0042930 | 0.0469943 | Paired amphipathic helix protein Sin3a |
| P09467 | -2.0219103 | 0.4961051 | 7.273827 | -4.075568 | 0.0043480 | 0.0473449 | Fructose-1,6-bisphosphatase 1 |
| Q9BVC6 | -1.6329609 | 0.4385611 | 9.420359 | -3.723452 | 0.0043822 | 0.0473476 | Transmembrane protein 109 |
| P56199 | -1.2716373 | 0.3331933 | 8.643688 | -3.816515 | 0.0044299 | 0.0473476 | Integrin alpha-1 |
| P50991 | -1.7929240 | 0.4666148 | 8.464244 | -3.842407 | 0.0044330 | 0.0473476 | T-complex protein 1 subunit delta |
| P51665 | -1.2958979 | 0.3474655 | 9.297522 | -3.729573 | 0.0044403 | 0.0473476 | 26S proteasome non-ATPase regulatory subunit 7 |
| Q6IC98 | -1.1710653 | 0.3080453 | 8.655845 | -3.801601 | 0.0045182 | 0.0478405 | GRAM domain-containing protein 4 |
| O43143 | -0.9916403 | 0.2678535 | 9.420359 | -3.702173 | 0.0045330 | 0.0478405 | Pre-mRNA-splicing factor ATP-dependent RNA helicase DHX15 |
| Q6ZWT7 | -2.6237461 | 0.5678695 | 5.420359 | -4.620333 | 0.0046816 | 0.0489641 | Lysophospholipid acyltransferase 2 |
| Q96FJ2 | 1.5408850 | 0.4077907 | 8.645240 | 3.778617 | 0.0046870 | 0.0489641 | Dynein light chain 2, cytoplasmic |
| Q6P587 | 5.5394450 | 1.3247288 | 6.582444 | 4.181569 | 0.0047308 | 0.0491316 | Acylpyruvase FAHD1, mitochondrial |
| P04843 | -2.0344244 | 0.5402204 | 8.671925 | -3.765915 | 0.0047508 | 0.0491316 | Dolichyl-diphosphooligosaccharide–protein glycosyltransferase subunit 1 |
| Q53FA7 | 1.8348852 | 0.4967783 | 9.200370 | 3.693570 | 0.0047852 | 0.0492400 | Quinone oxidoreductase PIG3 |
| P23434 | 1.1390966 | 0.3110180 | 9.420359 | 3.662478 | 0.0048291 | 0.0493121 | Glycine cleavage system H protein, mitochondrial |
| Q6P1N0 | -1.7290768 | 0.4259682 | 6.982437 | -4.059169 | 0.0048402 | 0.0493121 | Coiled-coil and C2 domain-containing protein 1A |
| Q8NBF2 | -1.1613746 | 0.3170052 | 9.347444 | -3.663582 | 0.0048840 | 0.0495134 | NHL repeat-containing protein 2 |
| A8MTT3 | -3.4393264 | 0.8546736 | 7.072356 | -4.024140 | 0.0049270 | 0.0497049 | Protein CEBPZOS |
| P80723 | -1.6306847 | 0.4256214 | 8.027411 | -3.831303 | 0.0049765 | 0.0499596 | Brain acid soluble protein 1 |
4.4 Evaluate results contrast \(\log_2 FC_{V-A}^R\)
4.4.1 Volcano-plot
volcanoRight <- ggplot(rowData(pe[["proteinRobust"]])$"tissueV + locationR:tissueV",
aes(x = logFC, y = -log10(pval), color = adjPval < 0.05)) +
geom_point(cex = 2.5) +
scale_color_manual(values = alpha(c("black", "red"), 0.5)) + theme_minimal()
volcanoRight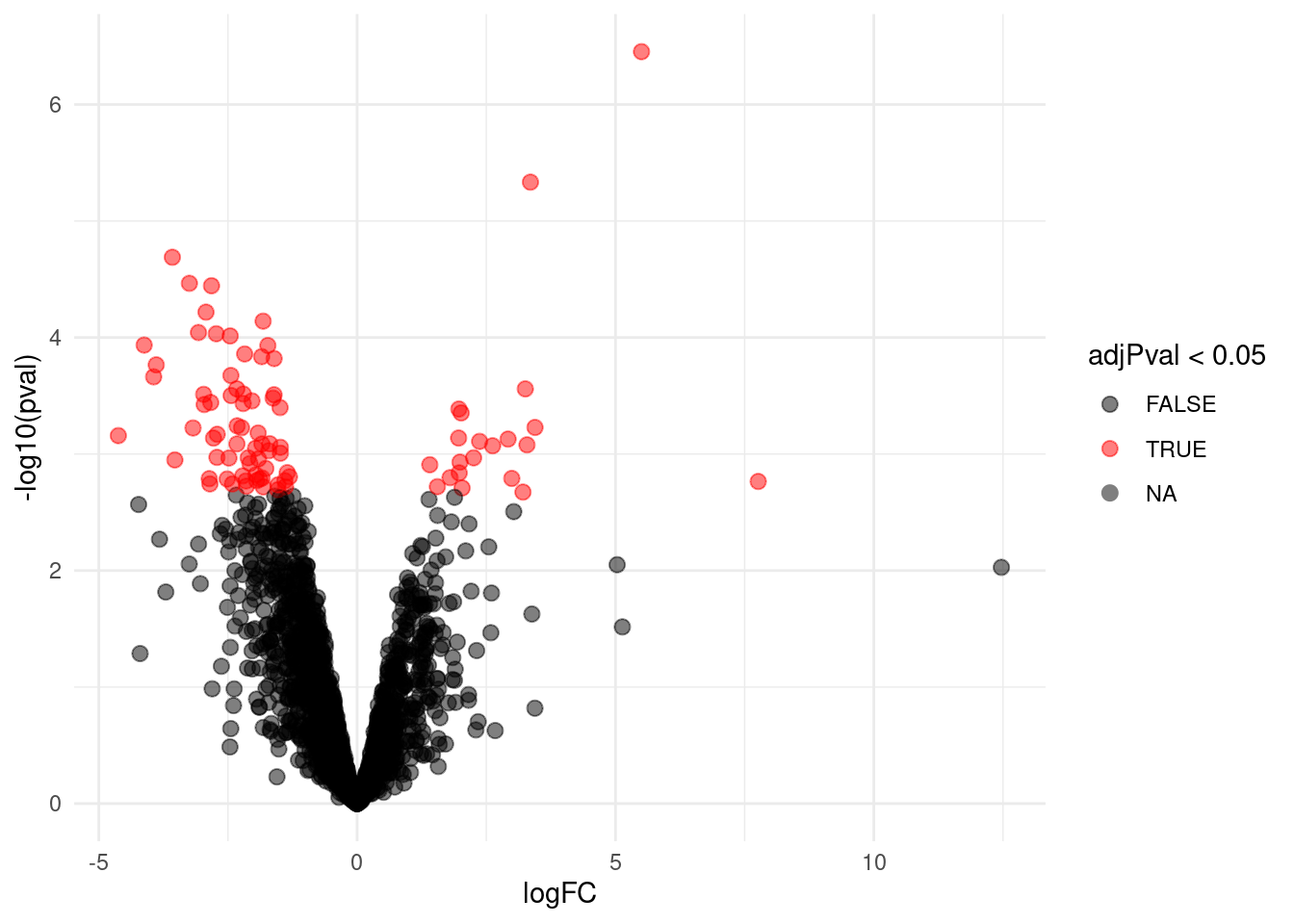
4.4.2 Heatmap
We first select the names of the proteins that were declared signficant.
sigNamesRight <- rowData(pe[["proteinRobust"]])$"tissueV + locationR:tissueV" %>%
rownames_to_column("proteinRobust") %>%
filter(adjPval<0.05) %>%
pull(proteinRobust)
heatmap(assay(pe[["proteinRobust"]])[sigNamesRight, ])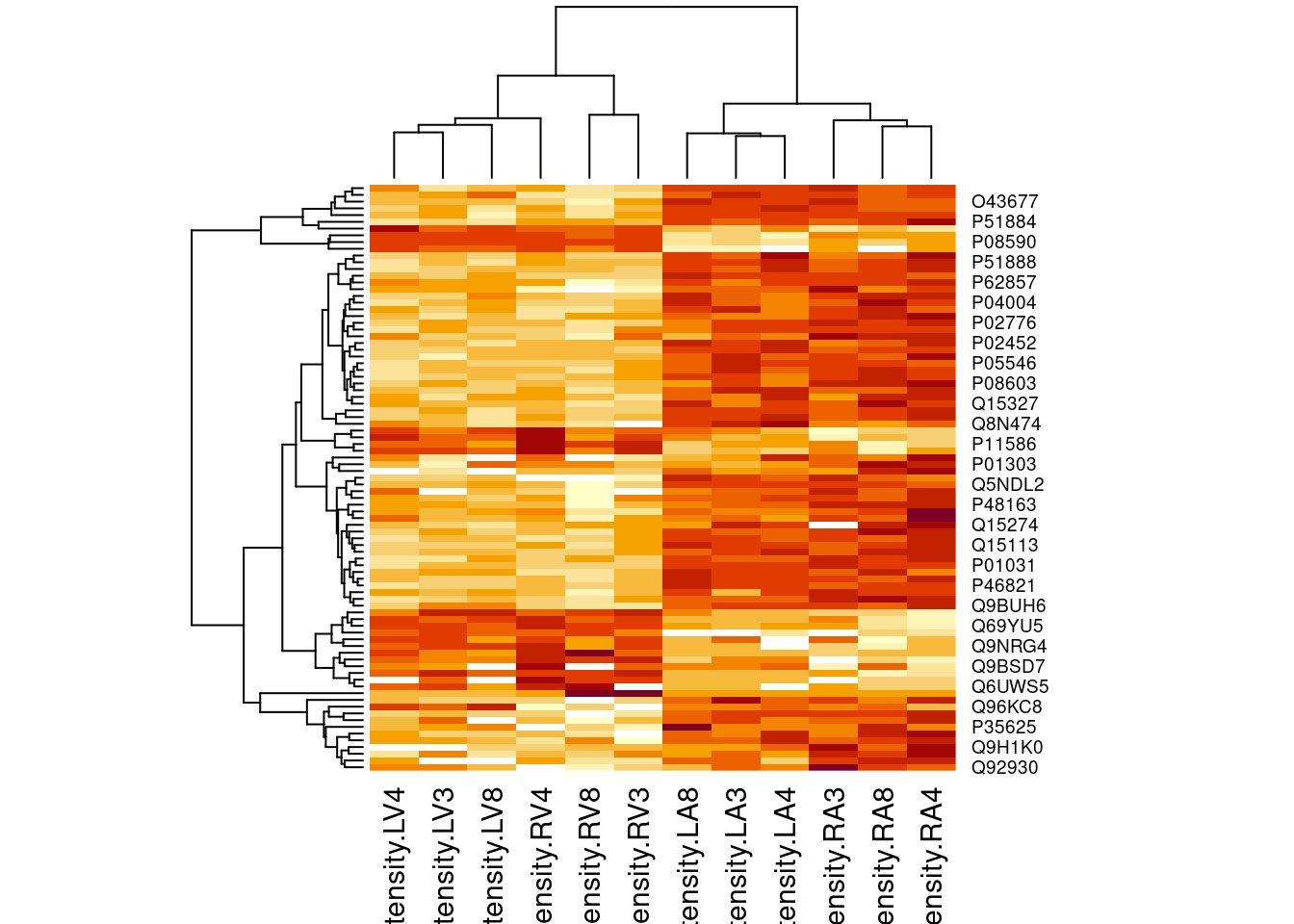
There are 87 proteins significantly differentially expressed at the 5% FDR level.
rowData(pe[["proteinRobust"]])$"tissueV + locationR:tissueV" %>%
cbind(.,rowData(pe[["proteinRobust"]])$Protein.names) %>%
na.exclude %>%
filter(adjPval<0.05) %>%
arrange(pval) %>%
knitr::kable(.)| logFC | se | df | t | pval | adjPval | rowData(pe[[“proteinRobust”]])$Protein.names | |
|---|---|---|---|---|---|---|---|
| P08590 | 5.503838 | 0.4386062 | 9.420359 | 12.548473 | 0.0000003 | 0.0006953 | Myosin light chain 3 |
| P06858 | 3.354674 | 0.3563885 | 9.302479 | 9.412969 | 0.0000047 | 0.0047476 | Lipoprotein lipase |
| Q9ULD0 | -3.575906 | 0.3771803 | 7.420359 | -9.480629 | 0.0000208 | 0.0141249 | 2-oxoglutarate dehydrogenase-like, mitochondrial |
| P35442 | -3.246460 | 0.4090718 | 8.420359 | -7.936162 | 0.0000346 | 0.0148139 | Thrombospondin-2 |
| P02776 | -2.818363 | 0.3445433 | 8.029185 | -8.179999 | 0.0000364 | 0.0148139 | Platelet factor 4;Platelet factor 4, short form |
| P21810 | -2.924783 | 0.4096541 | 8.810008 | -7.139639 | 0.0000605 | 0.0199120 | Biglycan |
| O75368 | -1.819326 | 0.2727814 | 9.342472 | -6.669538 | 0.0000769 | 0.0199120 | SH3 domain-binding glutamic acid-rich-like protein |
| A6NMZ7 | -3.070848 | 0.4717832 | 9.420359 | -6.509023 | 0.0000897 | 0.0199120 | Collagen alpha-6(VI) chain |
| P54652 | -2.726681 | 0.4132258 | 9.131756 | -6.598526 | 0.0000930 | 0.0199120 | Heat shock-related 70 kDa protein 2 |
| Q6UWY5 | -2.456874 | 0.3756784 | 9.166472 | -6.539833 | 0.0000979 | 0.0199120 | Olfactomedin-like protein 1 |
| Q06828 | -4.122091 | 0.6362258 | 9.022216 | -6.478974 | 0.0001130 | 0.0208866 | Fibromodulin |
| P05546 | -1.727800 | 0.2753193 | 9.324873 | -6.275623 | 0.0001246 | 0.0211179 | Heparin cofactor 2 |
| P28066 | -2.176824 | 0.3408032 | 8.782994 | -6.387335 | 0.0001415 | 0.0212787 | Proteasome subunit alpha type-5 |
| P29622 | -1.847919 | 0.3039016 | 9.420359 | -6.080651 | 0.0001521 | 0.0212787 | Kallistatin |
| P46821 | -1.606539 | 0.2664151 | 9.420359 | -6.030208 | 0.0001621 | 0.0212787 | Microtubule-associated protein 1B;MAP1B heavy chain;MAP1 light chain LC1 |
| P13533 | -3.888317 | 0.6374638 | 9.151585 | -6.099666 | 0.0001674 | 0.0212787 | Myosin-6 |
| Q9UGT4 | -2.443145 | 0.4157640 | 9.245036 | -5.876278 | 0.0002124 | 0.0240607 | Sushi domain-containing protein 2 |
| P35625 | -3.938312 | 0.6166712 | 7.993494 | -6.386405 | 0.0002129 | 0.0240607 | Metalloproteinase inhibitor 3 |
| Q69YU5 | 3.254738 | 0.5563163 | 8.748634 | 5.850517 | 0.0002718 | 0.0263699 | Uncharacterized protein C12orf73 |
| Q6PCB0 | -2.328852 | 0.4050843 | 8.987253 | -5.749055 | 0.0002781 | 0.0263699 | von Willebrand factor A domain-containing protein 1 |
| O43677 | -2.969964 | 0.5074118 | 8.483808 | -5.853162 | 0.0003049 | 0.0263699 | NADH dehydrogenase [ubiquinone] 1 subunit C1, mitochondrial |
| Q15327 | -2.203003 | 0.3952242 | 9.289627 | -5.574058 | 0.0003083 | 0.0263699 | Ankyrin repeat domain-containing protein 1 |
| P60468 | -2.438022 | 0.4136717 | 8.275558 | -5.893616 | 0.0003199 | 0.0263699 | Protein transport protein Sec61 subunit beta |
| Q14764 | -1.608232 | 0.2926175 | 9.420359 | -5.496021 | 0.0003253 | 0.0263699 | Major vault protein |
| P01031 | -1.626524 | 0.2847080 | 8.544930 | -5.712955 | 0.0003512 | 0.0263699 | Complement C5;Complement C5 beta chain;Complement C5 alpha chain;C5a anaphylatoxin;Complement C5 alpha chain |
| Q9P2B2 | -2.035226 | 0.3747097 | 9.420359 | -5.431474 | 0.0003548 | 0.0263699 | Prostaglandin F2 receptor negative regulator |
| Q92930 | -2.833666 | 0.4683580 | 7.661957 | -6.050213 | 0.0003628 | 0.0263699 | Ras-related protein Rab-8B |
| P02775 | -2.205543 | 0.4084962 | 9.420359 | -5.399176 | 0.0003707 | 0.0263699 | Platelet basic protein;Connective tissue-activating peptide III;TC-2;Connective tissue-activating peptide III(1-81);Beta-thromboglobulin;Neutrophil-activating peptide 2(74);Neutrophil-activating peptide 2(73);Neutrophil-activating peptide 2;TC-1;Neutrophil-activating peptide 2(1-66);Neutrophil-activating peptide 2(1-63) |
| P48163 | -2.959729 | 0.4904610 | 7.625022 | -6.034587 | 0.0003760 | 0.0263699 | NADP-dependent malic enzyme |
| P12883 | 1.969100 | 0.3673074 | 9.234880 | 5.360905 | 0.0004177 | 0.0277097 | Myosin-7 |
| P48681 | -1.491288 | 0.2811556 | 9.417394 | -5.304137 | 0.0004223 | 0.0277097 | Nestin |
| P11586 | 2.013301 | 0.3551466 | 8.048875 | 5.668929 | 0.0004607 | 0.0292833 | C-1-tetrahydrofolate synthase, cytoplasmic;Methylenetetrahydrofolate dehydrogenase;Methenyltetrahydrofolate cyclohydrolase;Formyltetrahydrofolate synthetase;C-1-tetrahydrofolate synthase, cytoplasmic, N-terminally processed |
| P30711 | -2.321740 | 0.4494189 | 9.090577 | -5.166094 | 0.0005717 | 0.0337808 | Glutathione S-transferase theta-1 |
| Q9BZH6 | 3.442805 | 0.6418925 | 8.373591 | 5.363523 | 0.0005792 | 0.0337808 | WD repeat-containing protein 11 |
| Q9UHG2 | -3.175806 | 0.6110813 | 8.897312 | -5.197026 | 0.0005876 | 0.0337808 | ProSAAS;KEP;Big SAAS;Little SAAS;Big PEN-LEN;PEN;Little LEN;Big LEN |
| Q9H1K0 | -2.237374 | 0.3926326 | 7.420359 | -5.698390 | 0.0005979 | 0.0337808 | Rabenosyn-5 |
| P00748 | -1.915326 | 0.3805658 | 9.156315 | -5.032839 | 0.0006705 | 0.0352687 | Coagulation factor XII;Coagulation factor XIIa heavy chain;Beta-factor XIIa part 1;Coagulation factor XIIa light chain |
| Q5NDL2 | -2.708130 | 0.5263639 | 8.717721 | -5.144976 | 0.0006715 | 0.0352687 | EGF domain-specific O-linked N-acetylglucosamine transferase |
| O00180 | -4.621308 | 0.8843846 | 8.420359 | -5.225451 | 0.0006762 | 0.0352687 | Potassium channel subfamily K member 1 |
| P23142 | -2.779235 | 0.5096807 | 7.589046 | -5.452894 | 0.0007260 | 0.0355824 | Fibulin-1 |
| P10916 | 2.921175 | 0.5305670 | 7.420359 | 5.505761 | 0.0007386 | 0.0355824 | Myosin regulatory light chain 2, ventricular/cardiac muscle isoform |
| Q00G26 | 1.964123 | 0.3627114 | 7.603807 | 5.415112 | 0.0007530 | 0.0355824 | Perilipin-5 |
| Q09161 | 2.370716 | 0.3907735 | 6.213723 | 6.066728 | 0.0007988 | 0.0355824 | Nuclear cap-binding protein subunit 1 |
| Q9BW30 | -2.326597 | 0.4732990 | 9.052429 | -4.915702 | 0.0008155 | 0.0355824 | Tubulin polymerization-promoting protein family member 3 |
| Q5JPH6 | 3.287773 | 0.5205786 | 5.829289 | 6.315614 | 0.0008229 | 0.0355824 | Probable glutamate–tRNA ligase, mitochondrial |
| P30405 | -1.842938 | 0.3822973 | 9.420359 | -4.820693 | 0.0008311 | 0.0355824 | Peptidyl-prolyl cis-trans isomerase F, mitochondrial |
| P18428 | -1.690875 | 0.3476064 | 9.191871 | -4.864339 | 0.0008380 | 0.0355824 | Lipopolysaccharide-binding protein |
| Q9NRG4 | 2.622493 | 0.5186766 | 8.420359 | 5.056124 | 0.0008397 | 0.0355824 | N-lysine methyltransferase SMYD2 |
| Q9BUH6 | -1.485484 | 0.3091856 | 9.196292 | -4.804507 | 0.0009105 | 0.0376245 | Protein PAXX |
| P35052 | -1.965366 | 0.3813736 | 7.847326 | -5.153388 | 0.0009249 | 0.0376245 | Glypican-1;Secreted glypican-1 |
| O15061 | -1.710970 | 0.3622447 | 9.420359 | -4.723242 | 0.0009567 | 0.0381573 | Synemin |
| P08603 | -1.484314 | 0.3171820 | 9.420359 | -4.679692 | 0.0010193 | 0.0390854 | Complement factor H |
| Q96FN9 | -2.712489 | 0.5555777 | 8.420359 | -4.882287 | 0.0010532 | 0.0390854 | Probable D-tyrosyl-tRNA(Tyr) deacylase 2 |
| Q04760 | 2.254328 | 0.4719185 | 8.783089 | 4.776944 | 0.0010764 | 0.0390854 | Lactoylglutathione lyase |
| Q96KC8 | -2.480944 | 0.5072249 | 8.308217 | -4.891212 | 0.0010821 | 0.0390854 | DnaJ homolog subfamily C member 1 |
| Q99983 | -3.527403 | 0.6599123 | 6.949931 | -5.345260 | 0.0010949 | 0.0390854 | Osteomodulin |
| Q8N474 | -2.108536 | 0.4231404 | 7.950693 | -4.983065 | 0.0010953 | 0.0390854 | Secreted frizzled-related protein 1 |
| P04004 | -1.908509 | 0.4017669 | 8.785099 | -4.750289 | 0.0011159 | 0.0391329 | Vitronectin;Vitronectin V65 subunit;Vitronectin V10 subunit;Somatomedin-B |
| P24298 | 1.991397 | 0.4203117 | 8.641195 | 4.737905 | 0.0011878 | 0.0409503 | Alanine aminotransferase 1 |
| P62857 | -2.074531 | 0.4177502 | 7.713404 | -4.965961 | 0.0012237 | 0.0414819 | 40S ribosomal protein S28 |
| P23434 | 1.406476 | 0.3110180 | 9.420359 | 4.522170 | 0.0012847 | 0.0428390 | Glycine cleavage system H protein, mitochondrial |
| P24844 | -1.767461 | 0.3927048 | 9.365146 | -4.500736 | 0.0013459 | 0.0441553 | Myosin regulatory light polypeptide 9 |
| Q6PI78 | 1.975693 | 0.4394104 | 9.092174 | 4.496236 | 0.0014592 | 0.0453981 | Transmembrane protein 65 |
| Q15113 | -1.959899 | 0.4411390 | 9.239033 | -4.442814 | 0.0015161 | 0.0453981 | Procollagen C-endopeptidase enhancer 1 |
| P04275 | -1.352764 | 0.3015061 | 8.996018 | -4.486688 | 0.0015194 | 0.0453981 | von Willebrand factor;von Willebrand antigen 2 |
| P01611 | -2.213481 | 0.5026161 | 9.420359 | -4.403920 | 0.0015321 | 0.0453981 | Ig kappa chain V-I region Wes |
| P04350 | -2.861432 | 0.5740170 | 6.993038 | -4.984925 | 0.0015968 | 0.0453981 | Tubulin beta-4A chain |
| Q9BSD7 | 2.993219 | 0.6098849 | 7.206934 | 4.907844 | 0.0015982 | 0.0453981 | Cancer-related nucleoside-triphosphatase |
| Q86WV6 | -1.310297 | 0.2989446 | 9.275839 | -4.383075 | 0.0016403 | 0.0453981 | Stimulator of interferon genes protein |
| P51888 | -1.838614 | 0.3767487 | 7.212550 | -4.880213 | 0.0016469 | 0.0453981 | Prolargin |
| Q6UWS5 | 1.798858 | 0.3707513 | 7.292033 | 4.851926 | 0.0016505 | 0.0453981 | Protein PET117 homolog, mitochondrial |
| P08294 | -1.884866 | 0.4063094 | 8.014829 | -4.638993 | 0.0016602 | 0.0453981 | Extracellular superoxide dismutase [Cu-Zn] |
| Q9UL26 | 7.763006 | 1.7464060 | 8.874918 | 4.445132 | 0.0016679 | 0.0453981 | Ras-related protein Rab-22A |
| Q7L4S7 | -2.514619 | 0.5153140 | 7.153052 | -4.879781 | 0.0016871 | 0.0453981 | Protein ARMCX6 |
| P01303 | -2.151374 | 0.4876274 | 8.990424 | -4.411922 | 0.0016952 | 0.0453981 | Pro-neuropeptide Y;Neuropeptide Y;C-flanking peptide of NPY |
| P82663 | -1.944516 | 0.4476294 | 9.372918 | -4.344030 | 0.0016963 | 0.0453981 | 28S ribosomal protein S25, mitochondrial |
| P50453 | -1.398216 | 0.3190416 | 8.998589 | -4.382552 | 0.0017652 | 0.0465364 | Serpin B9 |
| Q9HCB6 | -2.412515 | 0.5446307 | 8.694096 | -4.429636 | 0.0017933 | 0.0465364 | Spondin-1 |
| Q8TDB4 | -2.849906 | 0.5515757 | 6.286039 | -5.166844 | 0.0018075 | 0.0465364 | Protein MGARP |
| Q9Y6X5 | -1.531154 | 0.3489548 | 8.705094 | -4.387829 | 0.0018970 | 0.0472687 | Bis(5-adenosyl)-triphosphatase ENPP4 |
| Q15274 | -2.142900 | 0.4702421 | 7.916098 | -4.557015 | 0.0019081 | 0.0472687 | Nicotinate-nucleotide pyrophosphorylase [carboxylating] |
| P02452 | -1.816507 | 0.4086422 | 8.345480 | -4.445226 | 0.0019402 | 0.0472687 | Collagen alpha-1(I) chain |
| Q6YN16 | 1.553070 | 0.3657250 | 9.420359 | 4.246550 | 0.0019426 | 0.0472687 | Hydroxysteroid dehydrogenase-like protein 2 |
| P61925 | 2.031117 | 0.4388273 | 7.546436 | 4.628512 | 0.0019722 | 0.0472687 | cAMP-dependent protein kinase inhibitor alpha |
| P51884 | -1.386041 | 0.3246504 | 9.210584 | -4.269336 | 0.0019753 | 0.0472687 | Lumican |
| P07360 | -1.535545 | 0.3560274 | 8.758820 | -4.312997 | 0.0020806 | 0.0492078 | Complement component C8 gamma chain |
| Q96MM6 | 3.209344 | 0.6644094 | 6.744197 | 4.830370 | 0.0021064 | 0.0492466 | Heat shock 70 kDa protein 12B |
4.5 Evaluate results average contrast \(\log_2 FC_{V-A}\)
4.5.1 Volcano-plot
volcanoAvg <- ggplot(rowData(pe[["proteinRobust"]])$"tissueV + 0.5 * locationR:tissueV",
aes(x = logFC, y = -log10(pval), color = adjPval < 0.05)) +
geom_point(cex = 2.5) +
scale_color_manual(values = alpha(c("black", "red"), 0.5)) + theme_minimal()
volcanoAvg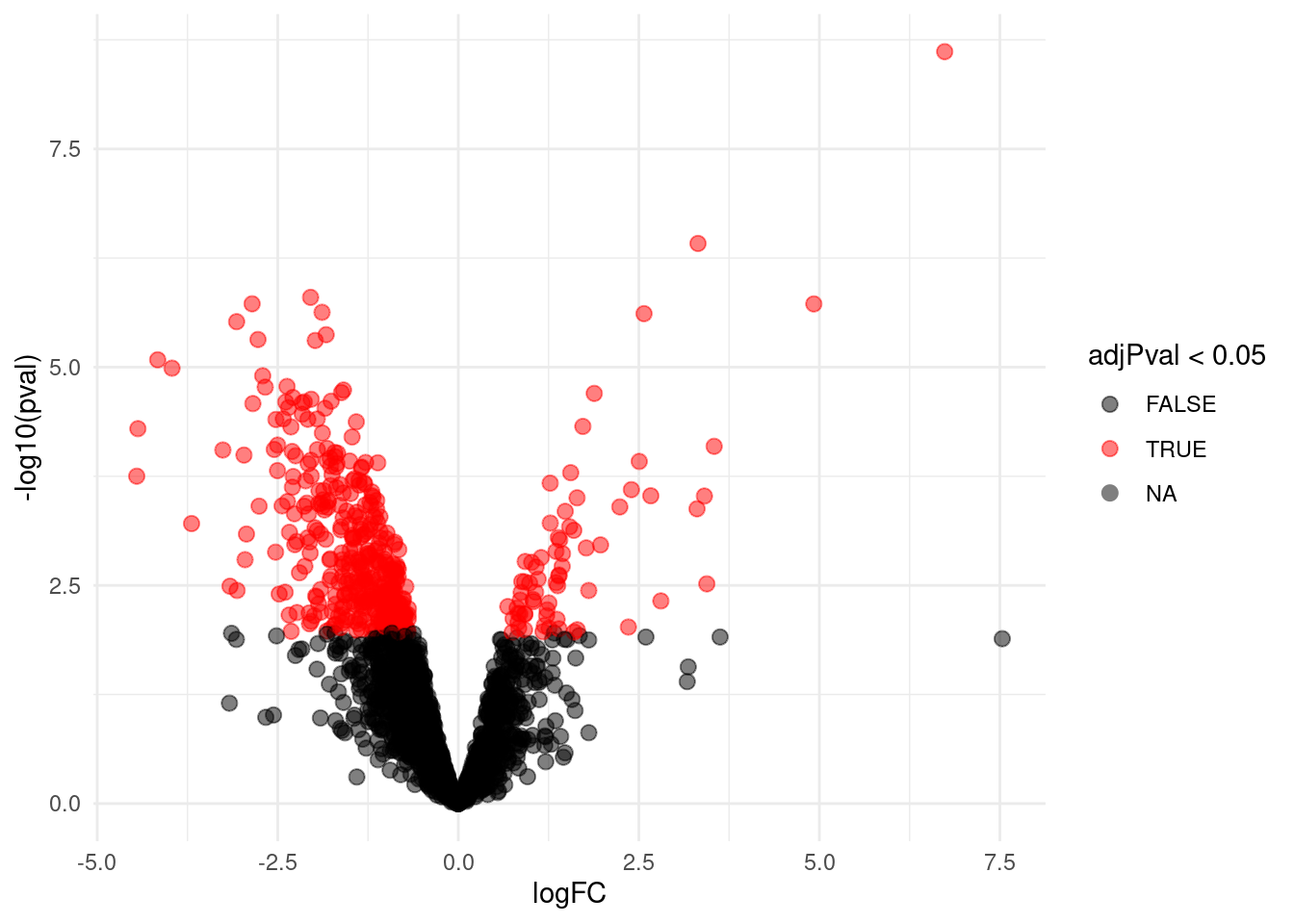
4.5.2 Heatmap
We first select the names of the proteins that were declared signficant.
sigNamesAvg <- rowData(pe[["proteinRobust"]])$"tissueV + 0.5 * locationR:tissueV" %>%
rownames_to_column("proteinRobust") %>%
filter(adjPval<0.05) %>%
pull(proteinRobust)
heatmap(assay(pe[["proteinRobust"]])[sigNamesAvg, ])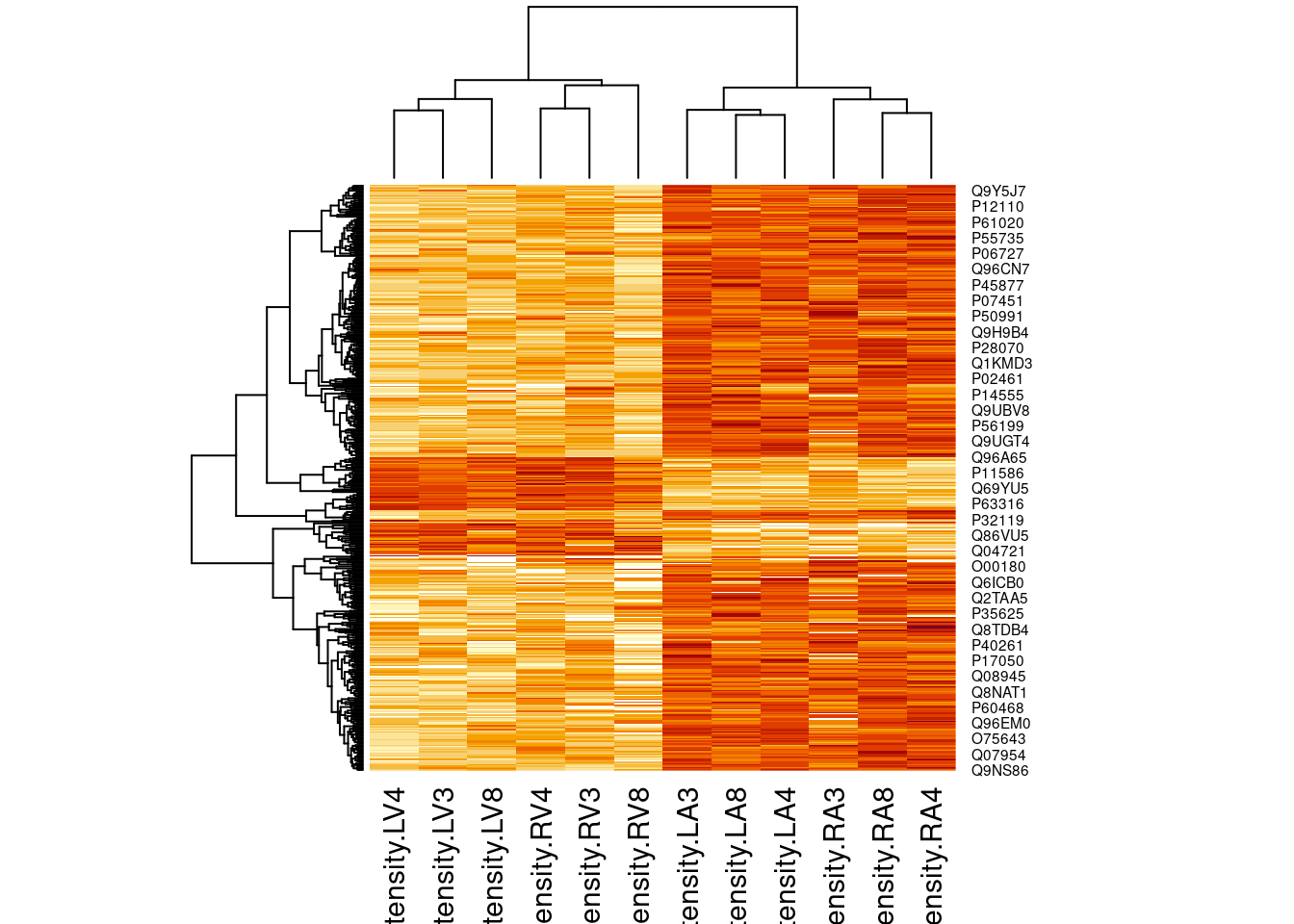
There are 445 proteins significantly differentially expressed at the 5% FDR level.
rowData(pe[["proteinRobust"]])$"tissueV + 0.5 * locationR:tissueV" %>%
cbind(.,rowData(pe[["proteinRobust"]])$Protein.names) %>%
na.exclude %>%
filter(adjPval<0.05) %>%
arrange(pval) %>%
knitr::kable(.)| logFC | se | df | t | pval | adjPval | rowData(pe[[“proteinRobust”]])$Protein.names | |
|---|---|---|---|---|---|---|---|
| P08590 | 6.7358945 | 0.3101414 | 9.420359 | 21.718785 | 0.0000000 | 0.0000047 | Myosin light chain 3 |
| P12883 | 3.3194914 | 0.2619286 | 9.234880 | 12.673268 | 0.0000004 | 0.0003850 | Myosin-7 |
| O75368 | -2.0454601 | 0.1935358 | 9.342472 | -10.568898 | 0.0000017 | 0.0007252 | SH3 domain-binding glutamic acid-rich-like protein |
| P10916 | 4.9236420 | 0.3670984 | 7.420359 | 13.412320 | 0.0000018 | 0.0007252 | Myosin regulatory light chain 2, ventricular/cardiac muscle isoform |
| Q6UWY5 | -2.8552584 | 0.2688443 | 9.166472 | -10.620493 | 0.0000019 | 0.0007252 | Olfactomedin-like protein 1 |
| P06858 | 2.5719330 | 0.2533166 | 9.302479 | 10.153038 | 0.0000024 | 0.0007252 | Lipoprotein lipase |
| P46821 | -1.8867094 | 0.1883840 | 9.420359 | -10.015234 | 0.0000025 | 0.0007252 | Microtubule-associated protein 1B;MAP1B heavy chain;MAP1 light chain LC1 |
| P21810 | -3.0696624 | 0.2942265 | 8.810008 | -10.432991 | 0.0000030 | 0.0007545 | Biglycan |
| P05546 | -1.8298412 | 0.1938785 | 9.324873 | -9.438084 | 0.0000045 | 0.0009443 | Heparin cofactor 2 |
| P35442 | -2.7737551 | 0.2705756 | 8.420359 | -10.251313 | 0.0000048 | 0.0009443 | Thrombospondin-2 |
| P29622 | -1.9806994 | 0.2148909 | 9.420359 | -9.217233 | 0.0000051 | 0.0009443 | Kallistatin |
| Q06828 | -4.1625547 | 0.4591032 | 9.022216 | -9.066708 | 0.0000079 | 0.0013397 | Fibromodulin |
| P13533 | -3.9643622 | 0.4544441 | 9.151585 | -8.723543 | 0.0000099 | 0.0015451 | Myosin-6 |
| Q8N474 | -2.7087798 | 0.2838348 | 7.950693 | -9.543507 | 0.0000125 | 0.0018211 | Secreted frizzled-related protein 1 |
| Q9UGT4 | -2.3718806 | 0.2916997 | 9.245036 | -8.131241 | 0.0000165 | 0.0020115 | Sushi domain-containing protein 2 |
| A6NMZ7 | -2.6735410 | 0.3336011 | 9.420359 | -8.014185 | 0.0000166 | 0.0020115 | Collagen alpha-6(VI) chain |
| O95865 | -1.5898713 | 0.2018470 | 9.420359 | -7.876616 | 0.0000192 | 0.0020115 | N(G),N(G)-dimethylarginine dimethylaminohydrolase 2 |
| Q14764 | -1.6175126 | 0.2069118 | 9.420359 | -7.817401 | 0.0000204 | 0.0020115 | Major vault protein |
| O94875-10 | 1.8826268 | 0.2322736 | 8.958125 | 8.105213 | 0.0000205 | 0.0020115 | Sorbin and SH3 domain-containing protein 2 |
| P02776 | -2.2899367 | 0.2620479 | 8.029185 | -8.738618 | 0.0000225 | 0.0020115 | Platelet factor 4;Platelet factor 4, short form |
| Q9P2B2 | -2.0373086 | 0.2649598 | 9.420359 | -7.689125 | 0.0000234 | 0.0020115 | Prostaglandin F2 receptor negative regulator |
| P02452 | -2.3897612 | 0.2863853 | 8.345480 | -8.344567 | 0.0000250 | 0.0020115 | Collagen alpha-1(I) chain |
| P24844 | -2.1318385 | 0.2783404 | 9.365146 | -7.659105 | 0.0000250 | 0.0020115 | Myosin regulatory light polypeptide 9 |
| P51884 | -1.7601864 | 0.2273949 | 9.210584 | -7.740660 | 0.0000252 | 0.0020115 | Lumican |
| O43677 | -2.8449373 | 0.3464798 | 8.483808 | -8.210977 | 0.0000256 | 0.0020115 | NADH dehydrogenase [ubiquinone] 1 subunit C1, mitochondrial |
| Q16647 | -2.1622988 | 0.2772049 | 9.081456 | -7.800362 | 0.0000257 | 0.0020115 | Prostacyclin synthase |
| Q15113 | -2.3511987 | 0.3094139 | 9.239033 | -7.598880 | 0.0000288 | 0.0021691 | Procollagen C-endopeptidase enhancer 1 |
| P18428 | -1.8452405 | 0.2433686 | 9.191871 | -7.582080 | 0.0000302 | 0.0021908 | Lipopolysaccharide-binding protein |
| P54652 | -2.1599833 | 0.2884317 | 9.131756 | -7.488717 | 0.0000345 | 0.0024215 | Heat shock-related 70 kDa protein 2 |
| Q9BW30 | -2.5255943 | 0.3406061 | 9.052429 | -7.415001 | 0.0000391 | 0.0024386 | Tubulin polymerization-promoting protein family member 3 |
| P02775 | -2.0838997 | 0.2888504 | 9.420359 | -7.214459 | 0.0000394 | 0.0024386 | Platelet basic protein;Connective tissue-activating peptide III;TC-2;Connective tissue-activating peptide III(1-81);Beta-thromboglobulin;Neutrophil-activating peptide 2(74);Neutrophil-activating peptide 2(73);Neutrophil-activating peptide 2;TC-1;Neutrophil-activating peptide 2(1-66);Neutrophil-activating peptide 2(1-63) |
| P51888 | -2.4233975 | 0.2735767 | 7.212550 | -8.858200 | 0.0000395 | 0.0024386 | Prolargin |
| P00748 | -1.9533971 | 0.2658345 | 9.156315 | -7.348170 | 0.0000396 | 0.0024386 | Coagulation factor XII;Coagulation factor XIIa heavy chain;Beta-factor XIIa part 1;Coagulation factor XIIa light chain |
| P48681 | -1.4123446 | 0.1987824 | 9.417394 | -7.104976 | 0.0000447 | 0.0026746 | Nestin |
| P08294 | -2.3175953 | 0.2941689 | 8.014829 | -7.878450 | 0.0000482 | 0.0027138 | Extracellular superoxide dismutase [Cu-Zn] |
| O00180 | -4.4369891 | 0.5849654 | 8.420359 | -7.585045 | 0.0000486 | 0.0027138 | Potassium channel subfamily K member 1 |
| P14854 | 1.7251878 | 0.2291779 | 8.484681 | 7.527724 | 0.0000494 | 0.0027138 | Cytochrome c oxidase subunit 6B1 |
| P02743 | -1.8823835 | 0.2732250 | 9.420359 | -6.889500 | 0.0000572 | 0.0030595 | Serum amyloid P-component;Serum amyloid P-component(1-203) |
| P01031 | -1.4711401 | 0.2044978 | 8.544930 | -7.193917 | 0.0000667 | 0.0034811 | Complement C5;Complement C5 beta chain;Complement C5 alpha chain;C5a anaphylatoxin;Complement C5 alpha chain |
| Q5NDL2 | -2.5014769 | 0.3591023 | 8.717721 | -6.965916 | 0.0000768 | 0.0038819 | EGF domain-specific O-linked N-acetylglucosamine transferase |
| Q92508 | 3.5430878 | 0.3997885 | 6.420359 | 8.862405 | 0.0000782 | 0.0038819 | Piezo-type mechanosensitive ion channel component 1 |
| Q53GQ0 | -1.8169289 | 0.2774067 | 9.410782 | -6.549695 | 0.0000858 | 0.0040140 | Very-long-chain 3-oxoacyl-CoA reductase |
| P05997 | -2.5486728 | 0.3787258 | 8.993385 | -6.729598 | 0.0000859 | 0.0040140 | Collagen alpha-2(V) chain |
| P23083 | -3.2601200 | 0.4241539 | 7.420359 | -7.686172 | 0.0000868 | 0.0040140 | Ig heavy chain V-I region V35 |
| P60468 | -1.9513827 | 0.2760385 | 8.275558 | -7.069240 | 0.0000891 | 0.0040256 | Protein transport protein Sec61 subunit beta |
| Q9ULL5-3 | -2.3006667 | 0.3165385 | 7.899339 | -7.268205 | 0.0000922 | 0.0040772 | Proline-rich protein 12 |
| Q8TBP6 | -1.7084465 | 0.2583618 | 9.032046 | -6.612613 | 0.0000963 | 0.0041153 | Solute carrier family 25 member 40 |
| P28066 | -1.6712466 | 0.2493234 | 8.782994 | -6.703127 | 0.0000989 | 0.0041153 | Proteasome subunit alpha type-5 |
| P35625 | -2.9679404 | 0.4160946 | 7.993494 | -7.132850 | 0.0000991 | 0.0041153 | Metalloproteinase inhibitor 3 |
| Q14195-2 | -2.2534620 | 0.3490857 | 9.283388 | -6.455327 | 0.0001022 | 0.0041571 | Dihydropyrimidinase-related protein 3 |
| P46060 | -1.7047689 | 0.2508453 | 8.420359 | -6.796096 | 0.0001089 | 0.0043393 | Ran GTPase-activating protein 1 |
| Q15327 | -1.7740861 | 0.2778695 | 9.289627 | -6.384601 | 0.0001109 | 0.0043393 | Ankyrin repeat domain-containing protein 1 |
| Q69YU5 | 2.5059630 | 0.3818797 | 8.748634 | 6.562178 | 0.0001180 | 0.0044197 | Uncharacterized protein C12orf73 |
| Q6PCB0 | -1.8163629 | 0.2815416 | 8.987253 | -6.451490 | 0.0001187 | 0.0044197 | von Willebrand factor A domain-containing protein 1 |
| Q9ULD0 | -2.0406330 | 0.2785661 | 7.420359 | -7.325489 | 0.0001195 | 0.0044197 | 2-oxoglutarate dehydrogenase-like, mitochondrial |
| P00325 | -1.5034066 | 0.2347510 | 9.037213 | -6.404261 | 0.0001224 | 0.0044456 | Alcohol dehydrogenase 1B |
| Q92604 | -2.0782169 | 0.3324288 | 9.352385 | -6.251615 | 0.0001267 | 0.0044705 | Acyl-CoA:lysophosphatidylglycerol acyltransferase 1 |
| P30405 | -1.6803249 | 0.2703250 | 9.420359 | -6.215944 | 0.0001284 | 0.0044705 | Peptidyl-prolyl cis-trans isomerase F, mitochondrial |
| P07451 | -1.2833190 | 0.2067250 | 9.420359 | -6.207855 | 0.0001297 | 0.0044705 | Carbonic anhydrase 3 |
| O14980 | -1.1146843 | 0.1804389 | 9.420359 | -6.177626 | 0.0001347 | 0.0045648 | Exportin-1 |
| P04004 | -1.7677440 | 0.2756173 | 8.785099 | -6.413763 | 0.0001371 | 0.0045725 | Vitronectin;Vitronectin V65 subunit;Vitronectin V10 subunit;Somatomedin-B |
| P36955 | -1.6933164 | 0.2541075 | 8.196767 | -6.663779 | 0.0001420 | 0.0046576 | Pigment epithelium-derived factor |
| P41240 | -1.3344948 | 0.2135594 | 9.050310 | -6.248821 | 0.0001463 | 0.0047120 | Tyrosine-protein kinase CSK |
| P04083 | -1.3421955 | 0.2200224 | 9.420359 | -6.100269 | 0.0001484 | 0.0047120 | Annexin A1 |
| P36021 | -2.5034435 | 0.3853203 | 8.420359 | -6.497046 | 0.0001506 | 0.0047120 | Monocarboxylate transporter 8 |
| Q6YN16 | 1.5567113 | 0.2586066 | 9.420359 | 6.019611 | 0.0001643 | 0.0050632 | Hydroxysteroid dehydrogenase-like protein 2 |
| Q8WWA0 | -4.4529908 | 0.6836913 | 8.156016 | -6.513160 | 0.0001704 | 0.0051332 | Intelectin-1 |
| Q8TBQ9 | -1.7482778 | 0.2790057 | 8.676104 | -6.266100 | 0.0001716 | 0.0051332 | Protein kish-A |
| Q9BXN1 | -2.2858485 | 0.3741823 | 9.012323 | -6.108916 | 0.0001763 | 0.0051428 | Asporin |
| Q15274 | -2.0357040 | 0.3082978 | 7.916098 | -6.603044 | 0.0001770 | 0.0051428 | Nicotinate-nucleotide pyrophosphorylase [carboxylating] |
| P04003 | -1.4359842 | 0.2384230 | 9.023047 | -6.022843 | 0.0001949 | 0.0055831 | C4b-binding protein alpha chain |
| Q7L4S7 | -2.1135247 | 0.3052384 | 7.153052 | -6.924177 | 0.0002050 | 0.0056579 | Protein ARMCX6 |
| Q9Y6X5 | -1.4593292 | 0.2392065 | 8.705094 | -6.100710 | 0.0002053 | 0.0056579 | Bis(5-adenosyl)-triphosphatase ENPP4 |
| O15230 | -1.3744676 | 0.2251448 | 8.689548 | -6.104816 | 0.0002058 | 0.0056579 | Laminin subunit alpha-5 |
| P14550 | -1.3048277 | 0.2179290 | 8.855961 | -5.987399 | 0.0002194 | 0.0058498 | Alcohol dehydrogenase [NADP(+)] |
| P23434 | 1.2727864 | 0.2199230 | 9.420359 | 5.787420 | 0.0002214 | 0.0058498 | Glycine cleavage system H protein, mitochondrial |
| Q9UBG0 | -1.6345146 | 0.2797211 | 9.225608 | -5.843373 | 0.0002233 | 0.0058498 | C-type mannose receptor 2 |
| Q07954 | -1.2959588 | 0.2245200 | 9.420359 | -5.772130 | 0.0002259 | 0.0058498 | Prolow-density lipoprotein receptor-related protein 1;Low-density lipoprotein receptor-related protein 1 85 kDa subunit;Low-density lipoprotein receptor-related protein 1 515 kDa subunit;Low-density lipoprotein receptor-related protein 1 intracellular domain |
| Q9BUF5 | -1.7682813 | 0.3045992 | 9.301065 | -5.805272 | 0.0002272 | 0.0058498 | Tubulin beta-6 chain |
| P49207 | -1.3710319 | 0.2330504 | 9.024427 | -5.882985 | 0.0002315 | 0.0058858 | 60S ribosomal protein L34 |
| P23142 | -2.2973847 | 0.3530386 | 7.589046 | -6.507461 | 0.0002348 | 0.0058969 | Fibulin-1 |
| P04196 | -1.6877176 | 0.2593507 | 7.506343 | -6.507472 | 0.0002462 | 0.0061031 | Histidine-rich glycoprotein |
| Q9NRG4 | 2.3970480 | 0.3961458 | 8.420359 | 6.050924 | 0.0002490 | 0.0061031 | N-lysine methyltransferase SMYD2 |
| Q9NZ01 | -1.8635396 | 0.3150317 | 8.685989 | -5.915403 | 0.0002583 | 0.0062151 | Very-long-chain enoyl-CoA reductase |
| O95980 | -1.9294212 | 0.3158816 | 8.199718 | -6.108051 | 0.0002597 | 0.0062151 | Reversion-inducing cysteine-rich protein with Kazal motifs |
| P12110 | -1.5935296 | 0.2644458 | 8.222232 | -6.025921 | 0.0002819 | 0.0065643 | Collagen alpha-2(VI) chain |
| P04275 | -1.1968998 | 0.2087610 | 8.996018 | -5.733349 | 0.0002826 | 0.0065643 | von Willebrand factor;von Willebrand antigen 2 |
| O75828 | -1.4912698 | 0.2663989 | 9.420359 | -5.597882 | 0.0002840 | 0.0065643 | Carbonyl reductase [NADPH] 3 |
| Q9BZH6 | 2.6654497 | 0.4489766 | 8.373591 | 5.936723 | 0.0002907 | 0.0066332 | WD repeat-containing protein 11 |
| Q5JPH6 | 3.4079519 | 0.4438849 | 5.829289 | 7.677558 | 0.0002935 | 0.0066332 | Probable glutamate–tRNA ligase, mitochondrial |
| P07357 | -1.1909118 | 0.2136116 | 9.262894 | -5.575127 | 0.0003111 | 0.0069535 | Complement component C8 alpha chain |
| P24298 | 1.6449487 | 0.2854239 | 8.641195 | 5.763178 | 0.0003168 | 0.0070046 | Alanine aminotransferase 1 |
| Q86WV6 | -1.1787564 | 0.2127534 | 9.275839 | -5.540483 | 0.0003240 | 0.0070857 | Stimulator of interferon genes protein |
| P62760 | -1.8748180 | 0.3370001 | 9.159766 | -5.563257 | 0.0003290 | 0.0071182 | Visinin-like protein 1 |
| P30711 | -1.7974916 | 0.3228953 | 9.090577 | -5.566793 | 0.0003365 | 0.0071979 | Glutathione S-transferase theta-1 |
| Q9UHG2 | -2.3658698 | 0.4209729 | 8.897312 | -5.620006 | 0.0003397 | 0.0071979 | ProSAAS;KEP;Big SAAS;Little SAAS;Big PEN-LEN;PEN;Little LEN;Big LEN |
| Q92681 | -1.8052155 | 0.3030168 | 7.932336 | -5.957477 | 0.0003505 | 0.0073112 | Regulatory solute carrier protein family 1 member 1 |
| P01024 | -1.1291448 | 0.2051109 | 9.157785 | -5.505046 | 0.0003552 | 0.0073112 | Complement C3;Complement C3 beta chain;C3-beta-c;Complement C3 alpha chain;C3a anaphylatoxin;Acylation stimulating protein;Complement C3b alpha chain;Complement C3c alpha chain fragment 1;Complement C3dg fragment;Complement C3g fragment;Complement C3d fragment;Complement C3f fragment;Complement C3c alpha chain fragment 2 |
| Q9HCB6 | -2.0954145 | 0.3709577 | 8.694096 | -5.648661 | 0.0003565 | 0.0073112 | Spondin-1 |
| P12814 | -1.9016090 | 0.3272561 | 8.209855 | -5.810766 | 0.0003631 | 0.0073112 | Alpha-actinin-1 |
| P48163 | -1.9351889 | 0.3194213 | 7.625022 | -6.058422 | 0.0003665 | 0.0073112 | NADP-dependent malic enzyme |
| Q9UQ35 | -1.2247564 | 0.2262830 | 9.401099 | -5.412499 | 0.0003666 | 0.0073112 | Serine/arginine repetitive matrix protein 2 |
| Q53GG5-2 | -2.4399434 | 0.4284092 | 8.409844 | -5.695357 | 0.0003804 | 0.0073955 | PDZ and LIM domain protein 3 |
| Q5M9N0 | -2.7584485 | 0.4838290 | 8.385642 | -5.701288 | 0.0003818 | 0.0073955 | Coiled-coil domain-containing protein 158 |
| P25940 | -1.6429098 | 0.3016823 | 9.155619 | -5.445828 | 0.0003843 | 0.0073955 | Collagen alpha-3(V) chain |
| P07585 | -2.1323372 | 0.3939309 | 9.259753 | -5.412972 | 0.0003860 | 0.0073955 | Decorin |
| Q9Y3B4 | -1.2875533 | 0.2358882 | 9.068421 | -5.458321 | 0.0003909 | 0.0073955 | Splicing factor 3B subunit 6 |
| P19429 | 2.2365865 | 0.3894775 | 8.211191 | 5.742531 | 0.0003929 | 0.0073955 | Troponin I, cardiac muscle |
| P50453 | -1.2078225 | 0.2212061 | 8.998589 | -5.460168 | 0.0004006 | 0.0073955 | Serpin B9 |
| P49458 | -1.8786710 | 0.3049990 | 7.251835 | -6.159597 | 0.0004026 | 0.0073955 | Signal recognition particle 9 kDa protein |
| Q53T59 | -1.8080103 | 0.3124888 | 8.037638 | -5.785841 | 0.0004046 | 0.0073955 | HCLS1-binding protein 3 |
| Q9UNW9 | 3.3060326 | 0.6115675 | 9.142176 | 5.405834 | 0.0004072 | 0.0073955 | RNA-binding protein Nova-2 |
| P02747 | -1.8530999 | 0.2992016 | 7.043683 | -6.193483 | 0.0004371 | 0.0078686 | Complement C1q subcomponent subunit C |
| O94919 | -1.1297756 | 0.2120168 | 9.184104 | -5.328709 | 0.0004443 | 0.0078985 | Endonuclease domain-containing 1 protein |
| P01008 | -1.5453986 | 0.2930302 | 9.376569 | -5.273855 | 0.0004466 | 0.0078985 | Antithrombin-III |
| Q00G26 | 1.4810539 | 0.2531323 | 7.603807 | 5.850908 | 0.0004631 | 0.0081128 | Perilipin-5 |
| O00264 | -1.4077482 | 0.2648534 | 9.097380 | -5.315198 | 0.0004670 | 0.0081128 | Membrane-associated progesterone receptor component 1 |
| O60760 | -2.2693850 | 0.4113993 | 8.420359 | -5.516259 | 0.0004707 | 0.0081128 | Hematopoietic prostaglandin D synthase |
| Q6SZW1 | -2.0732831 | 0.3507879 | 7.420359 | -5.910361 | 0.0004764 | 0.0081432 | Sterile alpha and TIR motif-containing protein 1 |
| Q9ULC3 | -1.4313739 | 0.2673476 | 8.720682 | -5.353981 | 0.0005115 | 0.0086695 | Ras-related protein Rab-23 |
| Q9BS26 | -1.1794250 | 0.2286961 | 9.354832 | -5.157170 | 0.0005282 | 0.0088132 | Endoplasmic reticulum resident protein 44 |
| Q9BTV4 | -1.5944595 | 0.3074186 | 9.211817 | -5.186607 | 0.0005329 | 0.0088132 | Transmembrane protein 43 |
| P14543 | -1.3732462 | 0.2639990 | 9.153301 | -5.201711 | 0.0005330 | 0.0088132 | Nidogen-1 |
| Q8WZA9 | -1.0848657 | 0.2099434 | 9.231251 | -5.167419 | 0.0005434 | 0.0088703 | Immunity-related GTPase family Q protein |
| P08603 | -1.1479994 | 0.2242815 | 9.420359 | -5.118564 | 0.0005451 | 0.0088703 | Complement factor H |
| Q9UKR5 | -3.6937409 | 0.6993749 | 8.547992 | -5.281489 | 0.0005997 | 0.0096809 | Probable ergosterol biosynthetic protein 28 |
| Q2TAA5 | -1.1153927 | 0.2205888 | 9.234456 | -5.056435 | 0.0006323 | 0.0100698 | GDP-Man:Man(3)GlcNAc(2)-PP-Dol alpha-1,2-mannosyltransferase |
| P11586 | 1.2736427 | 0.2359841 | 8.048875 | 5.397156 | 0.0006350 | 0.0100698 | C-1-tetrahydrofolate synthase, cytoplasmic;Methylenetetrahydrofolate dehydrogenase;Methenyltetrahydrofolate cyclohydrolase;Formyltetrahydrofolate synthetase;C-1-tetrahydrofolate synthase, cytoplasmic, N-terminally processed |
| P13671 | -1.3576211 | 0.2715324 | 9.420359 | -4.999850 | 0.0006439 | 0.0100698 | Complement component C6 |
| Q8WY22 | -1.5074312 | 0.2863698 | 8.420359 | -5.263932 | 0.0006441 | 0.0100698 | BRI3-binding protein |
| Q9UJC5 | -1.2143605 | 0.2426818 | 9.380287 | -5.003921 | 0.0006485 | 0.0100698 | SH3 domain-binding glutamic acid-rich-like protein 2 |
| O14967 | -1.6233066 | 0.3133456 | 8.580509 | -5.180563 | 0.0006742 | 0.0103497 | Calmegin |
| Q6PI78 | 1.5422623 | 0.3059022 | 9.092174 | 5.041684 | 0.0006767 | 0.0103497 | Transmembrane protein 65 |
| P01042 | -1.9829048 | 0.3533886 | 7.306380 | -5.611117 | 0.0006944 | 0.0105398 | Kininogen-1;Kininogen-1 heavy chain;T-kinin;Bradykinin;Lysyl-bradykinin;Kininogen-1 light chain;Low molecular weight growth-promoting factor |
| Q8WWQ0 | -1.3446280 | 0.2737312 | 9.420359 | -4.912220 | 0.0007291 | 0.0109135 | PH-interacting protein |
| P20774 | -1.9551933 | 0.3711074 | 8.070059 | -5.268538 | 0.0007355 | 0.0109135 | Mimecan |
| Q9HAT2 | 1.5992357 | 0.3240018 | 9.271229 | 4.935885 | 0.0007390 | 0.0109135 | Sialate O-acetylesterase |
| Q92621 | -1.2226648 | 0.2494199 | 9.417394 | -4.902035 | 0.0007404 | 0.0109135 | Nuclear pore complex protein Nup205 |
| Q12996 | -1.6262833 | 0.2731239 | 6.493841 | -5.954380 | 0.0007514 | 0.0109948 | Cleavage stimulation factor subunit 3 |
| Q8TDB4 | -2.3381087 | 0.3865801 | 6.286039 | -6.048187 | 0.0007778 | 0.0112937 | Protein MGARP |
| Q16082 | -1.2629563 | 0.2554732 | 9.059375 | -4.943596 | 0.0007829 | 0.0112937 | Heat shock protein beta-2 |
| Q9BUH6 | -1.0828646 | 0.2209139 | 9.196292 | -4.901749 | 0.0007940 | 0.0113017 | Protein PAXX |
| P02461 | -2.9334098 | 0.5751845 | 8.416773 | -5.099946 | 0.0007947 | 0.0113017 | Collagen alpha-1(III) chain |
| Q14314 | -1.9054905 | 0.3930629 | 9.417394 | -4.847800 | 0.0008001 | 0.0113017 | Fibroleukin |
| P45877 | -0.9878277 | 0.2050408 | 9.420359 | -4.817713 | 0.0008347 | 0.0116661 | Peptidyl-prolyl cis-trans isomerase C |
| P35052 | -1.2972987 | 0.2480341 | 7.847326 | -5.230325 | 0.0008431 | 0.0116661 | Glypican-1;Secreted glypican-1 |
| P02790 | -1.3146905 | 0.2731216 | 9.406717 | -4.813571 | 0.0008431 | 0.0116661 | Hemopexin |
| P01034 | -1.4676678 | 0.2975753 | 8.859133 | -4.932090 | 0.0008496 | 0.0116764 | Cystatin-C |
| P50479 | -2.0822296 | 0.4098960 | 8.192026 | -5.079898 | 0.0008863 | 0.0120871 | PDZ and LIM domain protein 4 |
| P40261 | -1.4439118 | 0.3008934 | 9.273912 | -4.798749 | 0.0008965 | 0.0120871 | Nicotinamide N-methyltransferase |
| Q86VP6 | -1.3506010 | 0.2719850 | 8.552957 | -4.965719 | 0.0009013 | 0.0120871 | Cullin-associated NEDD8-dissociated protein 1 |
| Q86VU5 | 1.3843626 | 0.2906522 | 9.420359 | 4.762952 | 0.0009033 | 0.0120871 | Catechol O-methyltransferase domain-containing protein 1 |
| P28300 | -1.4644405 | 0.2835416 | 7.825199 | -5.164816 | 0.0009203 | 0.0122349 | Protein-lysine 6-oxidase |
| Q9HAV4 | -1.8358132 | 0.3527066 | 7.670496 | -5.204930 | 0.0009344 | 0.0123394 | Exportin-5 |
| O95183 | -1.4512639 | 0.3064853 | 9.420359 | -4.735183 | 0.0009403 | 0.0123394 | Vesicle-associated membrane protein 5 |
| P06727 | -1.0726685 | 0.2210343 | 8.840955 | -4.852951 | 0.0009521 | 0.0124060 | Apolipoprotein A-IV |
| Q9NRX4 | 1.4050508 | 0.2956184 | 9.271509 | 4.752920 | 0.0009578 | 0.0124060 | 14 kDa phosphohistidine phosphatase |
| Q15126 | -0.9960340 | 0.2111020 | 9.420359 | -4.718261 | 0.0009637 | 0.0124060 | Phosphomevalonate kinase |
| Q96H79 | -2.2370397 | 0.3829114 | 6.200924 | -5.842187 | 0.0009863 | 0.0126172 | Zinc finger CCCH-type antiviral protein 1-like |
| O43175 | -2.0597036 | 0.4346794 | 9.221484 | -4.738443 | 0.0009925 | 0.0126175 | D-3-phosphoglycerate dehydrogenase |
| Q15582 | -2.2645249 | 0.4552646 | 8.068084 | -4.974086 | 0.0010605 | 0.0133968 | Transforming growth factor-beta-induced protein ig-h3 |
| P13667 | -1.0953363 | 0.2331484 | 9.172218 | -4.698022 | 0.0010670 | 0.0133968 | Protein disulfide-isomerase A4 |
| O43143 | -0.8789725 | 0.1894011 | 9.420359 | -4.640801 | 0.0010789 | 0.0134630 | Pre-mRNA-splicing factor ATP-dependent RNA helicase DHX15 |
| Q09161 | 1.9700086 | 0.3456614 | 6.213723 | 5.699243 | 0.0011179 | 0.0138642 | Nuclear cap-binding protein subunit 1 |
| P34932 | -0.8861862 | 0.1914748 | 9.291753 | -4.628214 | 0.0011396 | 0.0140482 | Heat shock 70 kDa protein 4 |
| Q9UKX3 | 1.7720370 | 0.3833701 | 9.238422 | 4.622261 | 0.0011671 | 0.0143000 | Myosin-13 |
| Q9NY15 | -1.4385610 | 0.3138718 | 9.403688 | -4.583275 | 0.0011793 | 0.0143640 | Stabilin-1 |
| P46940 | -1.1105871 | 0.2420668 | 9.239820 | -4.587936 | 0.0012261 | 0.0148443 | Ras GTPase-activating-like protein IQGAP1 |
| P26447 | -1.4990741 | 0.3207602 | 8.758554 | -4.673504 | 0.0012510 | 0.0150561 | Protein S100-A4 |
| O60262 | -1.2559109 | 0.2689796 | 8.700017 | -4.669167 | 0.0012814 | 0.0153110 | Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-7 |
| O95445 | -2.5298428 | 0.5522077 | 9.101762 | -4.581324 | 0.0012872 | 0.0153110 | Apolipoprotein M |
| Q96CS3 | -1.1510089 | 0.2514113 | 9.091487 | -4.578191 | 0.0012968 | 0.0153355 | FAS-associated factor 2 |
| P00747 | -0.8238672 | 0.1828138 | 9.406119 | -4.506593 | 0.0013198 | 0.0154483 | Plasminogen;Plasmin heavy chain A;Activation peptide;Angiostatin;Plasmin heavy chain A, short form;Plasmin light chain B |
| Q96JB2 | -2.0512914 | 0.4358269 | 8.420359 | -4.706665 | 0.0013299 | 0.0154483 | Conserved oligomeric Golgi complex subunit 3 |
| Q6UWS5 | 1.3529266 | 0.2687275 | 7.292033 | 5.034567 | 0.0013327 | 0.0154483 | Protein PET117 homolog, mitochondrial |
| P27658 | -1.6133702 | 0.3588925 | 9.420359 | -4.495414 | 0.0013367 | 0.0154483 | Collagen alpha-1(VIII) chain;Vastatin |
| P01019 | -1.0239723 | 0.2255684 | 9.127320 | -4.539521 | 0.0013573 | 0.0155978 | Angiotensinogen;Angiotensin-1;Angiotensin-2;Angiotensin-3;Angiotensin-4;Angiotensin 1-9;Angiotensin 1-7;Angiotensin 1-5;Angiotensin 1-4 |
| Q9BXV9 | 1.4430515 | 0.3205010 | 9.289309 | 4.502487 | 0.0013702 | 0.0156569 | Uncharacterized protein C14orf142 |
| Q07507 | -1.2545210 | 0.2616825 | 7.928623 | -4.794058 | 0.0014005 | 0.0159141 | Dermatopontin |
| P09619 | -1.2391198 | 0.2758540 | 9.226613 | -4.491941 | 0.0014154 | 0.0159936 | Platelet-derived growth factor receptor beta |
| P02748 | -1.2229145 | 0.2693768 | 8.912977 | -4.539792 | 0.0014417 | 0.0162007 | Complement component C9;Complement component C9a;Complement component C9b |
| Q96LL9 | -1.2440322 | 0.2745771 | 8.810329 | -4.530722 | 0.0015041 | 0.0168091 | DnaJ homolog subfamily C member 30 |
| Q92930 | -1.4774889 | 0.3083082 | 7.661957 | -4.792246 | 0.0015442 | 0.0171433 | Ras-related protein Rab-8B |
| Q04721 | 1.1483233 | 0.2590956 | 9.213519 | 4.432045 | 0.0015508 | 0.0171433 | Neurogenic locus notch homolog protein 2;Notch 2 extracellular truncation;Notch 2 intracellular domain |
| Q99983 | -2.9528995 | 0.5885590 | 6.949931 | -5.017168 | 0.0015683 | 0.0171916 | Osteomodulin |
| Q9UQR1 | -1.7724910 | 0.3607923 | 7.233799 | -4.912773 | 0.0015721 | 0.0171916 | Zinc finger protein 148 |
| P03950 | -1.7791248 | 0.3848924 | 8.205145 | -4.622395 | 0.0015950 | 0.0173485 | Angiogenin |
| O95486 | -1.3819063 | 0.3118915 | 9.078202 | -4.430728 | 0.0016110 | 0.0174297 | Protein transport protein Sec24A |
| P54577 | -1.0184999 | 0.2299667 | 9.016679 | -4.428901 | 0.0016423 | 0.0176342 | Tyrosine–tRNA ligase, cytoplasmic;Tyrosine–tRNA ligase, cytoplasmic, N-terminally processed |
| P83916 | -1.6056296 | 0.3090985 | 6.420359 | -5.194557 | 0.0016473 | 0.0176342 | Chromobox protein homolog 1 |
| Q9UBV8 | -1.1380629 | 0.2569081 | 8.957298 | -4.429844 | 0.0016668 | 0.0177499 | Peflin |
| Q7Z3T8 | -1.3746348 | 0.3062168 | 8.586144 | -4.489090 | 0.0017016 | 0.0180268 | Zinc finger FYVE domain-containing protein 16 |
| P00352 | -1.1346512 | 0.2499228 | 8.251350 | -4.540007 | 0.0017550 | 0.0181316 | Retinal dehydrogenase 1 |
| P62857 | -1.3551185 | 0.2898757 | 7.713404 | -4.674826 | 0.0017560 | 0.0181316 | 40S ribosomal protein S28 |
| P15924 | 0.9266546 | 0.2145531 | 9.383746 | 4.318998 | 0.0017566 | 0.0181316 | Desmoplakin |
| P80723 | -1.5107829 | 0.3290664 | 8.027411 | -4.591119 | 0.0017602 | 0.0181316 | Brain acid soluble protein 1 |
| O15061 | -1.1041834 | 0.2561457 | 9.420359 | -4.310763 | 0.0017625 | 0.0181316 | Synemin |
| Q9H1E5 | -1.0067111 | 0.2335851 | 9.420359 | -4.309826 | 0.0017650 | 0.0181316 | Thioredoxin-related transmembrane protein 4 |
| O75489 | 1.0163873 | 0.2360292 | 9.420359 | 4.306192 | 0.0017747 | 0.0181398 | NADH dehydrogenase [ubiquinone] iron-sulfur protein 3, mitochondrial |
| P19447 | -1.5879548 | 0.3346186 | 7.420359 | -4.745567 | 0.0017864 | 0.0181672 | TFIIH basal transcription factor complex helicase XPB subunit |
| Q9NS69 | -1.0847694 | 0.2462880 | 8.795400 | -4.404475 | 0.0018070 | 0.0182856 | Mitochondrial import receptor subunit TOM22 homolog |
| Q96CX2 | -0.9045513 | 0.2112848 | 9.341232 | -4.281195 | 0.0018791 | 0.0188636 | BTB/POZ domain-containing protein KCTD12 |
| Q08945 | -1.3761500 | 0.3092502 | 8.420359 | -4.449956 | 0.0018856 | 0.0188636 | FACT complex subunit SSRP1 |
| Q9NS86 | -0.9960486 | 0.2309453 | 9.124467 | -4.312920 | 0.0018919 | 0.0188636 | LanC-like protein 2 |
| O15118 | -2.1247916 | 0.4204381 | 6.420359 | -5.053756 | 0.0019057 | 0.0189078 | Niemann-Pick C1 protein |
| Q9Y287 | -1.6105936 | 0.3777397 | 9.358076 | -4.263766 | 0.0019212 | 0.0189117 | Integral membrane protein 2B;BRI2, membrane form;BRI2 intracellular domain;BRI2C, soluble form;Bri23 peptide |
| Q9BYN0 | -1.2801043 | 0.2981602 | 9.171396 | -4.293345 | 0.0019246 | 0.0189117 | Sulfiredoxin-1 |
| P52907 | -0.8523732 | 0.2010137 | 9.420359 | -4.240373 | 0.0019610 | 0.0190197 | F-actin-capping protein subunit alpha-1 |
| Q9UBB5 | 1.4388020 | 0.2863146 | 6.420359 | 5.025248 | 0.0019633 | 0.0190197 | Methyl-CpG-binding domain protein 2 |
| P62328 | -1.3455146 | 0.2900067 | 7.519371 | -4.639599 | 0.0019637 | 0.0190197 | Thymosin beta-4;Hematopoietic system regulatory peptide |
| Q8N142 | 1.0768090 | 0.2532957 | 9.279649 | 4.251193 | 0.0019957 | 0.0192379 | Adenylosuccinate synthetase isozyme 1 |
| P82663 | -1.3399129 | 0.3170820 | 9.372918 | -4.225761 | 0.0020278 | 0.0194551 | 28S ribosomal protein S25, mitochondrial |
| Q14118 | -0.8487363 | 0.1972071 | 8.781740 | -4.303782 | 0.0020956 | 0.0200113 | Dystroglycan;Alpha-dystroglycan;Beta-dystroglycan |
| P50991 | -1.5101244 | 0.3463126 | 8.464244 | -4.360581 | 0.0021080 | 0.0200355 | T-complex protein 1 subunit delta |
| P31323 | -0.9232942 | 0.2193346 | 9.248862 | -4.209524 | 0.0021407 | 0.0202522 | cAMP-dependent protein kinase type II-beta regulatory subunit |
| Q8NAT1 | -0.8308927 | 0.1986691 | 9.360836 | -4.182296 | 0.0021724 | 0.0204572 | Protein O-linked-mannose beta-1,4-N-acetylglucosaminyltransferase 2 |
| Q07065 | -0.9488623 | 0.2235778 | 8.959731 | -4.243992 | 0.0021835 | 0.0204668 | Cytoskeleton-associated protein 4 |
| P61009 | -0.8882212 | 0.2066525 | 8.595403 | -4.298139 | 0.0022203 | 0.0207006 | Signal peptidase complex subunit 3 |
| A5D6W6 | -2.1994339 | 0.4937525 | 7.849139 | -4.454527 | 0.0022288 | 0.0207006 | Fat storage-inducing transmembrane protein 1 |
| P07384 | -0.8641607 | 0.2077173 | 9.301416 | -4.160273 | 0.0022778 | 0.0210594 | Calpain-1 catalytic subunit |
| Q6IC98 | -0.9057366 | 0.2125145 | 8.655845 | -4.262000 | 0.0023004 | 0.0211723 | GRAM domain-containing protein 4 |
| Q96AG4 | -1.0641384 | 0.2572887 | 9.349851 | -4.135970 | 0.0023374 | 0.0214156 | Leucine-rich repeat-containing protein 59 |
| P05455 | -0.8562036 | 0.2063184 | 9.171164 | -4.149914 | 0.0023855 | 0.0217457 | Lupus La protein |
| Q12988 | -1.0103210 | 0.2443755 | 9.255593 | -4.134297 | 0.0023948 | 0.0217457 | Heat shock protein beta-3 |
| Q04760 | 1.3958984 | 0.3318358 | 8.783089 | 4.206593 | 0.0024115 | 0.0218004 | Lactoylglutathione lyase |
| P50238 | -1.7621993 | 0.4214564 | 8.861059 | -4.181214 | 0.0024543 | 0.0220228 | Cysteine-rich protein 1 |
| P62277 | -1.3005300 | 0.3173121 | 9.380882 | -4.098583 | 0.0024578 | 0.0220228 | 40S ribosomal protein S13 |
| P61925 | 1.3913131 | 0.3131368 | 7.546436 | 4.443148 | 0.0024919 | 0.0222302 | cAMP-dependent protein kinase inhibitor alpha |
| Q9H1K0 | -1.6022771 | 0.3584229 | 7.420359 | -4.470353 | 0.0025098 | 0.0222484 | Rabenosyn-5 |
| Q8IYI6 | -0.9330152 | 0.2287206 | 9.409497 | -4.079280 | 0.0025158 | 0.0222484 | Exocyst complex component 8 |
| P13796 | -0.8819572 | 0.2143660 | 9.129666 | -4.114258 | 0.0025417 | 0.0223806 | Plastin-2 |
| P05543 | -1.1801143 | 0.2649281 | 7.348700 | -4.454470 | 0.0026223 | 0.0229058 | Thyroxine-binding globulin |
| Q9UK22 | -1.3974278 | 0.2938770 | 6.412167 | -4.755145 | 0.0026280 | 0.0229058 | F-box only protein 2 |
| Q96A65 | -0.8321129 | 0.2055547 | 9.417394 | -4.048133 | 0.0026352 | 0.0229058 | Exocyst complex component 4 |
| P62745 | -1.1929691 | 0.2879947 | 8.729889 | -4.142330 | 0.0026838 | 0.0231598 | Rho-related GTP-binding protein RhoB |
| P58546 | -1.2963604 | 0.3163198 | 8.948856 | -4.098259 | 0.0027162 | 0.0231598 | Myotrophin |
| P24311 | 1.1056319 | 0.2745010 | 9.420359 | 4.027789 | 0.0027178 | 0.0231598 | Cytochrome c oxidase subunit 7B, mitochondrial |
| Q96FN9 | -1.7766077 | 0.4243295 | 8.420359 | -4.186859 | 0.0027240 | 0.0231598 | Probable D-tyrosyl-tRNA(Tyr) deacylase 2 |
| O00625 | -1.1028338 | 0.2740332 | 9.420359 | -4.024453 | 0.0027318 | 0.0231598 | Pirin |
| Q9Y490 | -0.8597753 | 0.2118978 | 9.188565 | -4.057500 | 0.0027327 | 0.0231598 | Talin-1 |
| O75348 | -1.2891767 | 0.3206941 | 9.420359 | -4.019958 | 0.0027510 | 0.0232176 | V-type proton ATPase subunit G 1 |
| P04843 | -1.5241453 | 0.3692851 | 8.671925 | -4.127286 | 0.0027829 | 0.0233905 | Dolichyl-diphosphooligosaccharide–protein glycosyltransferase subunit 1 |
| Q9UL18 | -1.3184619 | 0.3107203 | 7.913528 | -4.243243 | 0.0028955 | 0.0240366 | Protein argonaute-1 |
| Q8N5M1 | -1.2833584 | 0.3185222 | 9.122947 | -4.029103 | 0.0028956 | 0.0240366 | ATP synthase mitochondrial F1 complex assembly factor 2 |
| Q6P1N0 | -1.6508051 | 0.3697603 | 6.982437 | -4.464527 | 0.0029383 | 0.0240366 | Coiled-coil and C2 domain-containing protein 1A |
| Q96GK7 | -1.2557512 | 0.2967899 | 7.910446 | -4.231112 | 0.0029459 | 0.0240366 | Fumarylacetoacetate hydrolase domain-containing protein 2A |
| Q9NQZ5 | 1.3550021 | 0.3376320 | 9.152040 | 4.013252 | 0.0029470 | 0.0240366 | StAR-related lipid transfer protein 7, mitochondrial |
| Q9NNX1 | 3.4412993 | 0.7408907 | 6.420359 | 4.644813 | 0.0029547 | 0.0240366 | Tuftelin |
| P63316 | 0.8794535 | 0.2214963 | 9.420359 | 3.970512 | 0.0029707 | 0.0240366 | Troponin C, slow skeletal and cardiac muscles |
| Q15125 | -1.5522594 | 0.3856163 | 9.031289 | -4.025399 | 0.0029725 | 0.0240366 | 3-beta-hydroxysteroid-Delta(8),Delta(7)-isomerase |
| Q9Y4W6 | 0.9189304 | 0.2304862 | 9.296575 | 3.986921 | 0.0029732 | 0.0240366 | AFG3-like protein 2 |
| Q9HB40 | -1.4736038 | 0.3162446 | 6.361260 | -4.659697 | 0.0029780 | 0.0240366 | Retinoid-inducible serine carboxypeptidase |
| Q3ZCW2 | -1.3933792 | 0.3111768 | 6.883416 | -4.477773 | 0.0029961 | 0.0240872 | Galectin-related protein |
| Q96C86 | -0.8529077 | 0.2132782 | 9.149198 | -3.999038 | 0.0030135 | 0.0241315 | m7GpppX diphosphatase |
| Q9UKS6 | 0.9882506 | 0.2462779 | 8.991910 | 4.012745 | 0.0030568 | 0.0243623 | Protein kinase C and casein kinase substrate in neurons protein 3 |
| P00492 | -0.9642312 | 0.2405131 | 9.002876 | -4.009058 | 0.0030662 | 0.0243623 | Hypoxanthine-guanine phosphoribosyltransferase |
| Q00577 | -1.0249972 | 0.2562123 | 8.997651 | -4.000578 | 0.0031094 | 0.0246086 | Transcriptional activator protein Pur-alpha |
| P14555 | -3.1616763 | 0.7143843 | 6.916563 | -4.425736 | 0.0031505 | 0.0248378 | Phospholipase A2, membrane associated |
| Q53FA7 | 1.3758827 | 0.3477833 | 9.200370 | 3.956150 | 0.0031824 | 0.0249922 | Quinone oxidoreductase PIG3 |
| P09467 | -1.4304382 | 0.3334940 | 7.273827 | -4.289247 | 0.0033074 | 0.0258738 | Fructose-1,6-bisphosphatase 1 |
| P25311 | -1.0782056 | 0.2680130 | 8.556242 | -4.022960 | 0.0033319 | 0.0259657 | Zinc-alpha-2-glycoprotein |
| Q9UHD9 | -0.8978150 | 0.2281566 | 9.081182 | -3.935083 | 0.0033718 | 0.0261766 | Ubiquilin-2 |
| P01303 | -1.3886725 | 0.3523281 | 8.990424 | -3.941419 | 0.0034057 | 0.0262520 | Pro-neuropeptide Y;Neuropeptide Y;C-flanking peptide of NPY |
| P50552 | -0.9233883 | 0.2346915 | 9.036834 | -3.934477 | 0.0034073 | 0.0262520 | Vasodilator-stimulated phosphoprotein |
| Q7L4E1 | -1.9056994 | 0.4888519 | 9.203928 | -3.898317 | 0.0034763 | 0.0266529 | Protein FAM73B |
| O95810 | -0.7267426 | 0.1876918 | 9.391488 | -3.871999 | 0.0034856 | 0.0266529 | Serum deprivation-response protein |
| P53618 | -0.9076351 | 0.2349409 | 9.420359 | -3.863249 | 0.0035135 | 0.0267061 | Coatomer subunit beta |
| Q8IYU8 | -3.0629900 | 0.6825201 | 6.420359 | -4.487766 | 0.0035188 | 0.0267061 | Calcium uptake protein 2, mitochondrial |
| Q9BVC6 | -1.1958920 | 0.3101095 | 9.420359 | -3.856354 | 0.0035518 | 0.0268562 | Transmembrane protein 109 |
| Q96MM6 | 1.8061841 | 0.4138998 | 6.744197 | 4.363819 | 0.0036037 | 0.0271480 | Heat shock 70 kDa protein 12B |
| O15031 | -0.9094327 | 0.2339821 | 9.058686 | -3.886762 | 0.0036477 | 0.0273776 | Plexin-B2 |
| Q92575 | -0.9922287 | 0.2530061 | 8.755899 | -3.921758 | 0.0036939 | 0.0276226 | UBX domain-containing protein 4 |
| Q8IUX7 | -2.3978246 | 0.5007151 | 5.557874 | -4.788800 | 0.0037286 | 0.0277802 | Adipocyte enhancer-binding protein 1 |
| P28070 | -0.9263111 | 0.2288931 | 7.931934 | -4.046916 | 0.0037665 | 0.0278924 | Proteasome subunit beta type-4 |
| Q7KZF4 | -0.8794696 | 0.2306324 | 9.420359 | -3.813296 | 0.0038010 | 0.0278924 | Staphylococcal nuclease domain-containing protein 1 |
| P21980 | -0.9178411 | 0.2371165 | 8.976904 | -3.870844 | 0.0038023 | 0.0278924 | Protein-glutamine gamma-glutamyltransferase 2 |
| Q14258 | -0.9919848 | 0.2493945 | 8.251367 | -3.977573 | 0.0038293 | 0.0278924 | E3 ubiquitin/ISG15 ligase TRIM25 |
| A6NDG6 | 1.0711621 | 0.2790153 | 9.171051 | 3.839080 | 0.0038355 | 0.0278924 | Phosphoglycolate phosphatase |
| Q9Y2D4 | -1.5165389 | 0.3887140 | 8.722544 | -3.901426 | 0.0038359 | 0.0278924 | Exocyst complex component 6B |
| P27695 | -1.4919668 | 0.3919125 | 9.420359 | -3.806887 | 0.0038397 | 0.0278924 | DNA-(apurinic or apyrimidinic site) lyase;DNA-(apurinic or apyrimidinic site) lyase, mitochondrial |
| P43121 | -1.0780079 | 0.2743745 | 8.511897 | -3.928965 | 0.0038622 | 0.0279563 | Cell surface glycoprotein MUC18 |
| P14625 | -0.9671286 | 0.2543051 | 9.403167 | -3.803025 | 0.0038760 | 0.0279565 | Endoplasmin |
| Q9Y5U8 | -2.4800568 | 0.6509890 | 9.319580 | -3.809676 | 0.0038984 | 0.0280191 | Mitochondrial pyruvate carrier 1 |
| P10109 | 0.8708452 | 0.2279286 | 9.175781 | 3.820693 | 0.0039427 | 0.0282373 | Adrenodoxin, mitochondrial |
| E5RK69 | -1.1254137 | 0.2972634 | 9.420359 | -3.785914 | 0.0039690 | 0.0283264 | Annexin |
| Q15738 | -1.0818010 | 0.2866379 | 9.420359 | -3.774103 | 0.0040439 | 0.0285565 | Sterol-4-alpha-carboxylate 3-dehydrogenase, decarboxylating |
| P62312 | -1.5682756 | 0.4061040 | 8.734633 | -3.861759 | 0.0040596 | 0.0285565 | U6 snRNA-associated Sm-like protein LSm6 |
| Q01581 | -1.5736067 | 0.3821137 | 7.316536 | -4.118163 | 0.0040634 | 0.0285565 | Hydroxymethylglutaryl-CoA synthase, cytoplasmic |
| Q9BWJ5 | -1.1786521 | 0.3126119 | 9.420359 | -3.770336 | 0.0040681 | 0.0285565 | Splicing factor 3B subunit 5 |
| Q96PK6 | -0.9252023 | 0.2430058 | 9.118686 | -3.807326 | 0.0040715 | 0.0285565 | RNA-binding protein 14 |
| P04207 | -1.4064185 | 0.3683423 | 8.973244 | -3.818238 | 0.0041235 | 0.0288222 | Ig kappa chain V-III region CLL |
| Q13636 | -1.9750968 | 0.4490568 | 6.233646 | -4.398323 | 0.0041780 | 0.0290215 | Ras-related protein Rab-31 |
| P78406 | -1.6990379 | 0.3883613 | 6.300617 | -4.374890 | 0.0041812 | 0.0290215 | mRNA export factor |
| P31949 | -0.9710473 | 0.2584858 | 9.372327 | -3.756676 | 0.0041948 | 0.0290215 | Protein S100-A11;Protein S100-A11, N-terminally processed |
| P27169 | -1.0408608 | 0.2734080 | 8.938402 | -3.806987 | 0.0042252 | 0.0290356 | Serum paraoxonase/arylesterase 1 |
| P49773 | -0.9406572 | 0.2511758 | 9.420359 | -3.745015 | 0.0042347 | 0.0290356 | Histidine triad nucleotide-binding protein 1 |
| Q96EM0 | -1.2281296 | 0.3150239 | 8.300560 | -3.898529 | 0.0042397 | 0.0290356 | Trans-3-hydroxy-L-proline dehydratase |
| Q9BT09 | -1.2017444 | 0.3180938 | 9.105218 | -3.777957 | 0.0042723 | 0.0291493 | Protein canopy homolog 3 |
| P01699 | -1.9617461 | 0.4506972 | 6.301890 | -4.352692 | 0.0042850 | 0.0291493 | Ig lambda chain V-I region VOR |
| Q2TAY7 | -1.0380001 | 0.2752559 | 9.116096 | -3.771036 | 0.0043092 | 0.0292163 | WD40 repeat-containing protein SMU1;WD40 repeat-containing protein SMU1, N-terminally processed |
| Q04941 | -1.3354806 | 0.3557294 | 9.122186 | -3.754203 | 0.0044181 | 0.0297125 | Proteolipid protein 2 |
| P00488 | -1.1450813 | 0.3065552 | 9.274740 | -3.735319 | 0.0044192 | 0.0297125 | Coagulation factor XIII A chain |
| P01611 | -1.3210912 | 0.3554033 | 9.420359 | -3.717161 | 0.0044262 | 0.0297125 | Ig kappa chain V-I region Wes |
| Q9UNN8 | -1.2379487 | 0.3244063 | 8.506448 | -3.816044 | 0.0045663 | 0.0305523 | Endothelial protein C receptor |
| P07360 | -0.9508577 | 0.2519801 | 8.758820 | -3.773543 | 0.0046128 | 0.0307619 | Complement component C8 gamma chain |
| Q9BXF6 | -0.8964783 | 0.2431075 | 9.420359 | -3.687580 | 0.0046396 | 0.0308071 | Rab11 family-interacting protein 5 |
| Q5JUQ0 | 2.8054144 | 0.6808551 | 6.864054 | 4.120428 | 0.0046498 | 0.0308071 | Protein FAM78A |
| Q02818 | -1.1067621 | 0.2711064 | 6.995049 | -4.082390 | 0.0046826 | 0.0308706 | Nucleobindin-1 |
| Q9HBL0 | 1.0363941 | 0.2720098 | 8.420359 | 3.810135 | 0.0046939 | 0.0308706 | Tensin-1 |
| Q96KP1 | -1.0392328 | 0.2587608 | 7.267948 | -4.016191 | 0.0047050 | 0.0308706 | Exocyst complex component 2 |
| O75165 | -0.7578962 | 0.2049879 | 9.231691 | -3.697272 | 0.0047298 | 0.0309079 | DnaJ homolog subfamily C member 13 |
| P20042 | -0.8792944 | 0.2376566 | 9.197242 | -3.699852 | 0.0047410 | 0.0309079 | Eukaryotic translation initiation factor 2 subunit 2 |
| P46063 | -0.7705141 | 0.2085324 | 9.183164 | -3.694937 | 0.0047903 | 0.0309698 | ATP-dependent DNA helicase Q1 |
| Q8IVD9 | -0.9380220 | 0.2450845 | 8.198729 | -3.827341 | 0.0048105 | 0.0309698 | NudC domain-containing protein 3 |
| P10643 | -1.1503065 | 0.3113889 | 9.159735 | -3.694115 | 0.0048176 | 0.0309698 | Complement component C7 |
| P31689 | -1.4848105 | 0.3875317 | 8.155261 | -3.831456 | 0.0048300 | 0.0309698 | DnaJ homolog subfamily A member 1 |
| Q13011 | 0.8580498 | 0.2299097 | 8.838577 | 3.732116 | 0.0048338 | 0.0309698 | Delta(3,5)-Delta(2,4)-dienoyl-CoA isomerase, mitochondrial |
| Q04837 | -1.2706968 | 0.3386600 | 8.651174 | -3.752131 | 0.0048713 | 0.0309698 | Single-stranded DNA-binding protein, mitochondrial |
| Q9BU61 | -1.2937160 | 0.3537628 | 9.420359 | -3.657015 | 0.0048714 | 0.0309698 | NADH dehydrogenase [ubiquinone] 1 alpha subcomplex assembly factor 3 |
| Q9BXR6 | -1.3408193 | 0.3666219 | 9.417394 | -3.657227 | 0.0048723 | 0.0309698 | Complement factor H-related protein 5 |
| Q08357 | -1.2512681 | 0.3317594 | 8.465455 | -3.771613 | 0.0049202 | 0.0311310 | Sodium-dependent phosphate transporter 2 |
| Q53GG5 | 1.0396066 | 0.2802467 | 8.915226 | 3.709613 | 0.0049283 | 0.0311310 | PDZ and LIM domain protein 3 |
| Q9Y5U9 | -0.9688010 | 0.2537423 | 8.113937 | -3.818051 | 0.0049709 | 0.0313029 | Immediate early response 3-interacting protein 1 |
| Q8NDY3 | 1.2548368 | 0.3412954 | 9.066122 | 3.676689 | 0.0050383 | 0.0316295 | [Protein ADP-ribosylarginine] hydrolase-like protein 1 |
| Q9HCN8 | -0.8376149 | 0.2299564 | 9.325084 | -3.642495 | 0.0050707 | 0.0317347 | Stromal cell-derived factor 2-like protein 1 |
| Q9UBI9 | -1.9350473 | 0.4883540 | 7.231219 | -3.962387 | 0.0050999 | 0.0318195 | Headcase protein homolog |
| O75663 | -1.6975991 | 0.4618211 | 8.989551 | -3.675880 | 0.0051181 | 0.0318358 | TIP41-like protein |
| P00918 | -0.9493414 | 0.2613767 | 9.327391 | -3.632081 | 0.0051532 | 0.0318941 | Carbonic anhydrase 2 |
| Q8TDB6 | -0.9819803 | 0.2685382 | 9.102046 | -3.656762 | 0.0051629 | 0.0318941 | E3 ubiquitin-protein ligase DTX3L |
| Q9Y5J7 | -1.4394711 | 0.3977241 | 9.420359 | -3.619271 | 0.0051746 | 0.0318941 | Mitochondrial import inner membrane translocase subunit Tim9 |
| P61020 | -0.8349426 | 0.2271616 | 8.866958 | -3.675545 | 0.0052423 | 0.0321784 | Ras-related protein Rab-5B |
| O15121 | -1.3734454 | 0.3782388 | 9.228763 | -3.631159 | 0.0052523 | 0.0321784 | Sphingolipid delta(4)-desaturase DES1 |
| O00299 | -0.9153236 | 0.2528154 | 9.304824 | -3.620521 | 0.0052698 | 0.0321887 | Chloride intracellular channel protein 1 |
| Q8NBF2 | -0.8115466 | 0.2248622 | 9.347444 | -3.609084 | 0.0053267 | 0.0323921 | NHL repeat-containing protein 2 |
| P31994-2 | -0.7713913 | 0.2123017 | 9.122267 | -3.633466 | 0.0053350 | 0.0323921 | Low affinity immunoglobulin gamma Fc region receptor II-b |
| Q8WUM0 | -0.7928646 | 0.2201827 | 9.277093 | -3.600940 | 0.0054630 | 0.0330708 | Nuclear pore complex protein Nup133 |
| Q15493 | -1.5865116 | 0.4375846 | 9.005754 | -3.625611 | 0.0055174 | 0.0333009 | Regucalcin |
| P46777 | -1.0578875 | 0.2897786 | 8.757208 | -3.650676 | 0.0055626 | 0.0334533 | 60S ribosomal protein L5 |
| P00742 | -1.3361965 | 0.3268762 | 6.419136 | -4.087775 | 0.0055841 | 0.0334533 | Coagulation factor X;Factor X light chain;Factor X heavy chain;Activated factor Xa heavy chain |
| O00567 | -1.6066654 | 0.4489340 | 9.345669 | -3.578845 | 0.0055920 | 0.0334533 | Nucleolar protein 56 |
| P49756 | -0.8733625 | 0.2452468 | 9.420359 | -3.561158 | 0.0056804 | 0.0338825 | RNA-binding protein 25 |
| Q9H4A6 | -0.9863755 | 0.2771451 | 9.333154 | -3.559058 | 0.0057841 | 0.0344004 | Golgi phosphoprotein 3 |
| Q6PCE3 | -1.3068366 | 0.3405543 | 7.360778 | -3.837380 | 0.0058132 | 0.0344727 | Glucose 1,6-bisphosphate synthase |
| O15144 | -1.0266458 | 0.2845878 | 8.851301 | -3.607483 | 0.0058393 | 0.0345266 | Actin-related protein 2/3 complex subunit 2 |
| Q6P1L8 | 0.6856534 | 0.1923234 | 9.202756 | 3.565107 | 0.0058582 | 0.0345381 | 39S ribosomal protein L14, mitochondrial |
| P24752 | 0.8145171 | 0.2270837 | 8.915799 | 3.586858 | 0.0059577 | 0.0350232 | Acetyl-CoA acetyltransferase, mitochondrial |
| P06396 | -1.1557324 | 0.3175614 | 8.455087 | -3.639398 | 0.0060041 | 0.0351939 | Gelsolin |
| Q1KMD3 | -0.9588418 | 0.2691474 | 9.065422 | -3.562515 | 0.0060251 | 0.0352156 | Heterogeneous nuclear ribonucleoprotein U-like protein 2 |
| Q96PE7 | -1.1472298 | 0.3258058 | 9.420359 | -3.521208 | 0.0060579 | 0.0353060 | Methylmalonyl-CoA epimerase, mitochondrial |
| Q9HAN9 | 1.2305740 | 0.3230453 | 7.321157 | 3.809293 | 0.0060984 | 0.0354402 | Nicotinamide/nicotinic acid mononucleotide adenylyltransferase 1 |
| P56199 | -0.8817502 | 0.2451200 | 8.643688 | -3.597218 | 0.0061682 | 0.0356539 | Integrin alpha-1 |
| Q9Y646 | -0.6906706 | 0.1960589 | 9.294244 | -3.522770 | 0.0061702 | 0.0356539 | Carboxypeptidase Q |
| P62330 | -0.7931556 | 0.2242913 | 9.122775 | -3.536275 | 0.0062172 | 0.0358132 | ADP-ribosylation factor 6 |
| Q9UBS4 | -1.7470606 | 0.4930682 | 9.044784 | -3.543244 | 0.0062330 | 0.0358132 | DnaJ homolog subfamily B member 11 |
| P17050 | -0.9454418 | 0.2682555 | 9.156208 | -3.524408 | 0.0062990 | 0.0360905 | Alpha-N-acetylgalactosaminidase |
| Q9Y5Z7 | -1.9019574 | 0.4680155 | 6.116460 | -4.063877 | 0.0063592 | 0.0362794 | Host cell factor 2 |
| P01597 | -2.2333895 | 0.6327136 | 9.042973 | -3.529859 | 0.0063676 | 0.0362794 | Ig kappa chain V-I region DEE |
| P05062 | -2.0647894 | 0.5508210 | 7.417394 | -3.748567 | 0.0064626 | 0.0367177 | Fructose-bisphosphate aldolase B |
| P62191 | -1.0000517 | 0.2855948 | 9.201844 | -3.501645 | 0.0064811 | 0.0367200 | 26S protease regulatory subunit 4 |
| Q8TCJ2 | -0.8698436 | 0.2465217 | 8.921075 | -3.528467 | 0.0065189 | 0.0368316 | Dolichyl-diphosphooligosaccharide–protein glycosyltransferase subunit STT3B |
| Q9H9B4 | -0.7776690 | 0.2196810 | 8.758716 | -3.539992 | 0.0065917 | 0.0371397 | Sideroflexin-1 |
| P08754 | -1.0292580 | 0.2979223 | 9.420359 | -3.454787 | 0.0067445 | 0.0375627 | Guanine nucleotide-binding protein G(k) subunit alpha |
| P30050 | -1.1309945 | 0.3257840 | 9.240612 | -3.471608 | 0.0067562 | 0.0375627 | 60S ribosomal protein L12 |
| Q68DH5 | -0.9839434 | 0.2752828 | 8.344587 | -3.574301 | 0.0067644 | 0.0375627 | LMBR1 domain-containing protein 2 |
| Q14019 | -2.3417241 | 0.5964209 | 6.420359 | -3.926294 | 0.0067706 | 0.0375627 | Coactosin-like protein |
| P23284 | -0.7744408 | 0.2214611 | 8.982458 | -3.496960 | 0.0067761 | 0.0375627 | Peptidyl-prolyl cis-trans isomerase B |
| P60903 | -0.8289441 | 0.2386403 | 9.201691 | -3.473613 | 0.0067775 | 0.0375627 | Protein S100-A10 |
| O43920 | 0.8295850 | 0.2386795 | 9.164468 | 3.475728 | 0.0067962 | 0.0375640 | NADH dehydrogenase [ubiquinone] iron-sulfur protein 5 |
| Q9GZY4 | -1.6172207 | 0.4432059 | 7.775949 | -3.648916 | 0.0068292 | 0.0376440 | Cytochrome c oxidase assembly factor 1 homolog |
| P15848 | 0.9204565 | 0.2603319 | 8.570614 | 3.535704 | 0.0068679 | 0.0376645 | Arylsulfatase B |
| Q9HAV7 | 0.8532336 | 0.2459635 | 9.164544 | 3.468945 | 0.0068700 | 0.0376645 | GrpE protein homolog 1, mitochondrial |
| O14807 | -0.9860449 | 0.2846002 | 9.178081 | -3.464667 | 0.0069016 | 0.0377246 | Ras-related protein M-Ras |
| Q9Y623 | -1.9955095 | 0.5618523 | 8.399371 | -3.551662 | 0.0069240 | 0.0377246 | Myosin-4 |
| P27797 | -0.8755252 | 0.2546797 | 9.417394 | -3.437751 | 0.0069366 | 0.0377246 | Calreticulin |
| O75643 | -0.7286175 | 0.2121708 | 9.420359 | -3.434109 | 0.0069744 | 0.0378294 | U5 small nuclear ribonucleoprotein 200 kDa helicase |
| Q12907 | -1.6602075 | 0.4671559 | 8.313980 | -3.553862 | 0.0070158 | 0.0378649 | Vesicular integral-membrane protein VIP36 |
| O75190-3 | 1.2231318 | 0.3565719 | 9.420359 | 3.430253 | 0.0070182 | 0.0378649 | DnaJ homolog subfamily B member 6 |
| P17540 | 0.9216483 | 0.2487257 | 7.298674 | 3.705481 | 0.0070540 | 0.0379570 | Creatine kinase S-type, mitochondrial |
| O43290 | -0.7436611 | 0.2171716 | 9.420359 | -3.424301 | 0.0070863 | 0.0380306 | U4/U6.U5 tri-snRNP-associated protein 1 |
| P98160 | -0.6944147 | 0.2022592 | 9.245897 | -3.433291 | 0.0071780 | 0.0383683 | Basement membrane-specific heparan sulfate proteoglycan core protein;Endorepellin;LG3 peptide |
| Q16204 | -0.7607082 | 0.2190201 | 8.855932 | -3.473235 | 0.0071870 | 0.0383683 | Coiled-coil domain-containing protein 6 |
| P21281 | -1.3428216 | 0.3928941 | 9.377634 | -3.417769 | 0.0072093 | 0.0383866 | V-type proton ATPase subunit B, brain isoform |
| P30101 | -0.7361350 | 0.2161032 | 9.401877 | -3.406405 | 0.0073160 | 0.0388531 | Protein disulfide-isomerase A3 |
| Q6ZVF9 | -1.5316163 | 0.3710478 | 5.510860 | -4.127814 | 0.0073979 | 0.0391859 | G protein-regulated inducer of neurite outgrowth 3 |
| P41208 | -1.1048997 | 0.3226308 | 9.121806 | -3.424657 | 0.0074238 | 0.0392206 | Centrin-2 |
| O75533 | -0.7480960 | 0.2199021 | 9.281221 | -3.401950 | 0.0075073 | 0.0395594 | Splicing factor 3B subunit 1 |
| O43707 | -1.3709413 | 0.3951980 | 8.554624 | -3.468999 | 0.0076265 | 0.0400835 | Alpha-actinin-4 |
| P56192 | -1.0251690 | 0.2985340 | 8.830575 | -3.434010 | 0.0076742 | 0.0401467 | Methionine–tRNA ligase, cytoplasmic |
| P32119 | -0.8759792 | 0.2468160 | 7.897094 | -3.549119 | 0.0076787 | 0.0401467 | Peroxiredoxin-2 |
| P55735 | -0.9148870 | 0.2659790 | 8.745427 | -3.439697 | 0.0077172 | 0.0401467 | Protein SEC13 homolog |
| Q92805 | 1.3677629 | 0.3990676 | 8.858631 | 3.427396 | 0.0077177 | 0.0401467 | Golgin subfamily A member 1 |
| Q8NBJ5 | -1.0090957 | 0.2932132 | 8.713942 | -3.441508 | 0.0077372 | 0.0401467 | Procollagen galactosyltransferase 1 |
| P02749 | -0.9821379 | 0.2881999 | 8.996525 | -3.407836 | 0.0077806 | 0.0402690 | Beta-2-glycoprotein 1 |
| Q9UK41 | -1.5562781 | 0.4606623 | 9.268585 | -3.378349 | 0.0078138 | 0.0403382 | Vacuolar protein sorting-associated protein 28 homolog |
| Q5VUM1 | 0.7587056 | 0.2245024 | 9.213910 | 3.379499 | 0.0078652 | 0.0405009 | Succinate dehydrogenase assembly factor 4, mitochondrial |
| Q96G03 | -0.8738839 | 0.2603156 | 9.420359 | -3.357017 | 0.0079060 | 0.0406079 | Phosphoglucomutase-2 |
| P43686 | -1.2985795 | 0.3875167 | 9.398102 | -3.351029 | 0.0080098 | 0.0409929 | 26S protease regulatory subunit 6B |
| Q6ICB0 | -2.0382992 | 0.5471065 | 6.695413 | -3.725599 | 0.0080212 | 0.0409929 | Desumoylating isopeptidase 1 |
| P11021 | -0.6930674 | 0.2048069 | 9.016794 | -3.384004 | 0.0080539 | 0.0410209 | 78 kDa glucose-regulated protein |
| Q9BVG4 | -1.4282890 | 0.4270292 | 9.419676 | -3.344711 | 0.0080670 | 0.0410209 | Protein PBDC1 |
| Q6Y288 | -1.0690665 | 0.3184078 | 9.258387 | -3.357539 | 0.0080932 | 0.0410515 | Beta-1,3-glucosyltransferase |
| Q9UBQ0 | -1.1043560 | 0.3293908 | 9.239664 | -3.352722 | 0.0081795 | 0.0413860 | Vacuolar protein sorting-associated protein 29 |
| P61026 | -0.8659400 | 0.2443342 | 7.602181 | -3.544081 | 0.0082280 | 0.0415280 | Ras-related protein Rab-10 |
| O60568 | -1.0635431 | 0.3203369 | 9.420359 | -3.320077 | 0.0083972 | 0.0422680 | Procollagen-lysine,2-oxoglutarate 5-dioxygenase 3 |
| Q14011 | -1.6060873 | 0.4676141 | 8.287145 | -3.434642 | 0.0084318 | 0.0422680 | Cold-inducible RNA-binding protein |
| Q5T447 | -1.0258262 | 0.3044486 | 8.870208 | -3.369456 | 0.0084370 | 0.0422680 | E3 ubiquitin-protein ligase HECTD3 |
| P17174 | 0.8269867 | 0.2485633 | 9.191878 | 3.327066 | 0.0085865 | 0.0428091 | Aspartate aminotransferase, cytoplasmic |
| Q96LD4 | -2.0603233 | 0.5325761 | 5.888034 | -3.868599 | 0.0085871 | 0.0428091 | Tripartite motif-containing protein 47 |
| Q9BX97 | -1.3847012 | 0.3974417 | 7.758076 | -3.484036 | 0.0086779 | 0.0431559 | Plasmalemma vesicle-associated protein |
| P51692 | -1.6950637 | 0.5096371 | 9.100029 | -3.326021 | 0.0087208 | 0.0432637 | Signal transducer and activator of transcription 5B |
| Q9HCJ6 | -0.8677868 | 0.2633701 | 9.417394 | -3.294932 | 0.0087533 | 0.0433192 | Synaptic vesicle membrane protein VAT-1 homolog-like |
| Q9BSH5 | -0.8065712 | 0.2419806 | 8.978305 | -3.333206 | 0.0087837 | 0.0433643 | Haloacid dehalogenase-like hydrolase domain-containing protein 3 |
| P29992 | -1.0018710 | 0.2959582 | 8.436243 | -3.385178 | 0.0088516 | 0.0435938 | Guanine nucleotide-binding protein subunit alpha-11 |
| A4D2B0 | -1.7140517 | 0.5055811 | 8.367083 | -3.390261 | 0.0088896 | 0.0436751 | Metallo-beta-lactamase domain-containing protein 1 |
| P16455 | -0.7988054 | 0.2358633 | 8.351627 | -3.386731 | 0.0089616 | 0.0439229 | Methylated-DNA–protein-cysteine methyltransferase |
| P05387 | -0.7447264 | 0.2265778 | 9.261556 | -3.286846 | 0.0090698 | 0.0443459 | 60S acidic ribosomal protein P2 |
| P16083 | -1.0028849 | 0.2977294 | 8.408267 | -3.368444 | 0.0091246 | 0.0444935 | Ribosyldihydronicotinamide dehydrogenase [quinone] |
| I3L505 | 1.1922390 | 0.3578892 | 8.742159 | 3.331308 | 0.0091437 | 0.0444935 | Acyl carrier protein |
| O60927 | -0.9472893 | 0.2671994 | 7.072548 | -3.545252 | 0.0092452 | 0.0448802 | Protein phosphatase 1 regulatory subunit 11 |
| Q13425 | -0.7615654 | 0.2331007 | 9.307500 | -3.267109 | 0.0093039 | 0.0450325 | Beta-2-syntrophin |
| Q8N392 | 2.3552996 | 0.6366348 | 6.272040 | 3.699609 | 0.0093209 | 0.0450325 | Rho GTPase-activating protein 18 |
| Q96CN7 | -1.0144660 | 0.3102486 | 9.191660 | -3.269849 | 0.0094174 | 0.0453910 | Isochorismatase domain-containing protein 1 |
| P01621 | -1.1890923 | 0.3622603 | 8.991047 | -3.282425 | 0.0095052 | 0.0457056 | Ig kappa chain V-III region NG9 |
| Q9UMR3 | -1.6328959 | 0.4714071 | 7.420359 | -3.463876 | 0.0095749 | 0.0459325 | T-box transcription factor TBX20 |
| Q9NX08 | -0.7274661 | 0.2225081 | 8.998441 | -3.269391 | 0.0096944 | 0.0463004 | COMM domain-containing protein 8 |
| O43592 | -0.8026366 | 0.2477266 | 9.327348 | -3.240009 | 0.0096971 | 0.0463004 | Exportin-T |
| Q13825 | 1.3948092 | 0.4330846 | 9.420359 | 3.220639 | 0.0098829 | 0.0470010 | Methylglutaconyl-CoA hydratase, mitochondrial |
| O95159 | -1.7663091 | 0.5267649 | 8.061146 | -3.353126 | 0.0099239 | 0.0470010 | Zinc finger protein-like 1 |
| O60503 | 1.2572380 | 0.3432644 | 6.207277 | 3.662594 | 0.0099335 | 0.0470010 | Adenylate cyclase type 9 |
| P35754 | 0.8371729 | 0.2590987 | 9.256564 | 3.231097 | 0.0099363 | 0.0470010 | Glutaredoxin-1 |
| Q6UXG3 | 1.6474617 | 0.5114802 | 9.266008 | 3.220969 | 0.0100882 | 0.0476088 | CMRF35-like molecule 9 |
| Q6B0K9 | -1.3448839 | 0.3998344 | 7.854249 | -3.363603 | 0.0101471 | 0.0477759 | Hemoglobin subunit mu |
| P03915 | -1.1507202 | 0.3495468 | 8.417376 | -3.292035 | 0.0102419 | 0.0480857 | NADH-ubiquinone oxidoreductase chain 5 |
| P34947 | -1.4080509 | 0.4396529 | 9.361568 | -3.202642 | 0.0102602 | 0.0480857 | G protein-coupled receptor kinase 5 |
| Q13541 | 0.9175109 | 0.2871743 | 9.420359 | 3.194961 | 0.0103090 | 0.0482035 | Eukaryotic translation initiation factor 4E-binding protein 1 |
| O14656 | -1.8183830 | 0.5541802 | 8.420359 | -3.281212 | 0.0104087 | 0.0485580 | Torsin-1A |
| P05060 | -2.3136323 | 0.6771858 | 7.313959 | -3.416540 | 0.0104618 | 0.0486940 | Secretogranin-1;PE-11;GAWK peptide;CCB peptide |
| P14174 | -0.8418646 | 0.2617826 | 9.024325 | -3.215892 | 0.0105221 | 0.0488630 | Macrophage migration inhibitory factor |
| P19474 | -1.1365988 | 0.3417059 | 7.930483 | -3.326248 | 0.0105730 | 0.0489788 | E3 ubiquitin-protein ligase TRIM21 |
| O75436 | -0.9929865 | 0.3067959 | 8.751304 | -3.236636 | 0.0105952 | 0.0489788 | Vacuolar protein sorting-associated protein 26A |
| Q08AG7 | -1.1984106 | 0.3496044 | 7.161376 | -3.427905 | 0.0106361 | 0.0490564 | Mitotic-spindle organizing protein 1 |
| Q8IWU2 | -0.7035098 | 0.2137465 | 8.159580 | -3.291327 | 0.0107028 | 0.0491840 | Serine/threonine-protein kinase LMTK2 |
| Q8IWB7 | 1.6050991 | 0.4816712 | 7.808945 | 3.332354 | 0.0107121 | 0.0491840 | WD repeat and FYVE domain-containing protein 1 |
| Q0VAK6 | 1.1733962 | 0.3606850 | 8.492777 | 3.253244 | 0.0107427 | 0.0492132 | Leiomodin-3 |
| P07196 | -3.1402185 | 0.8492355 | 5.757771 | -3.697701 | 0.0108927 | 0.0497880 | Neurofilament light polypeptide |
4.6 Interaction
4.6.1 Volcano-plot
volcanoInt <- ggplot(rowData(pe[["proteinRobust"]])$"locationR:tissueV",
aes(x = logFC, y = -log10(pval), color = adjPval < 0.05)) +
geom_point(cex = 2.5) +
scale_color_manual(values = alpha(c("black", "red"), 0.5)) + theme_minimal()
volcanoInt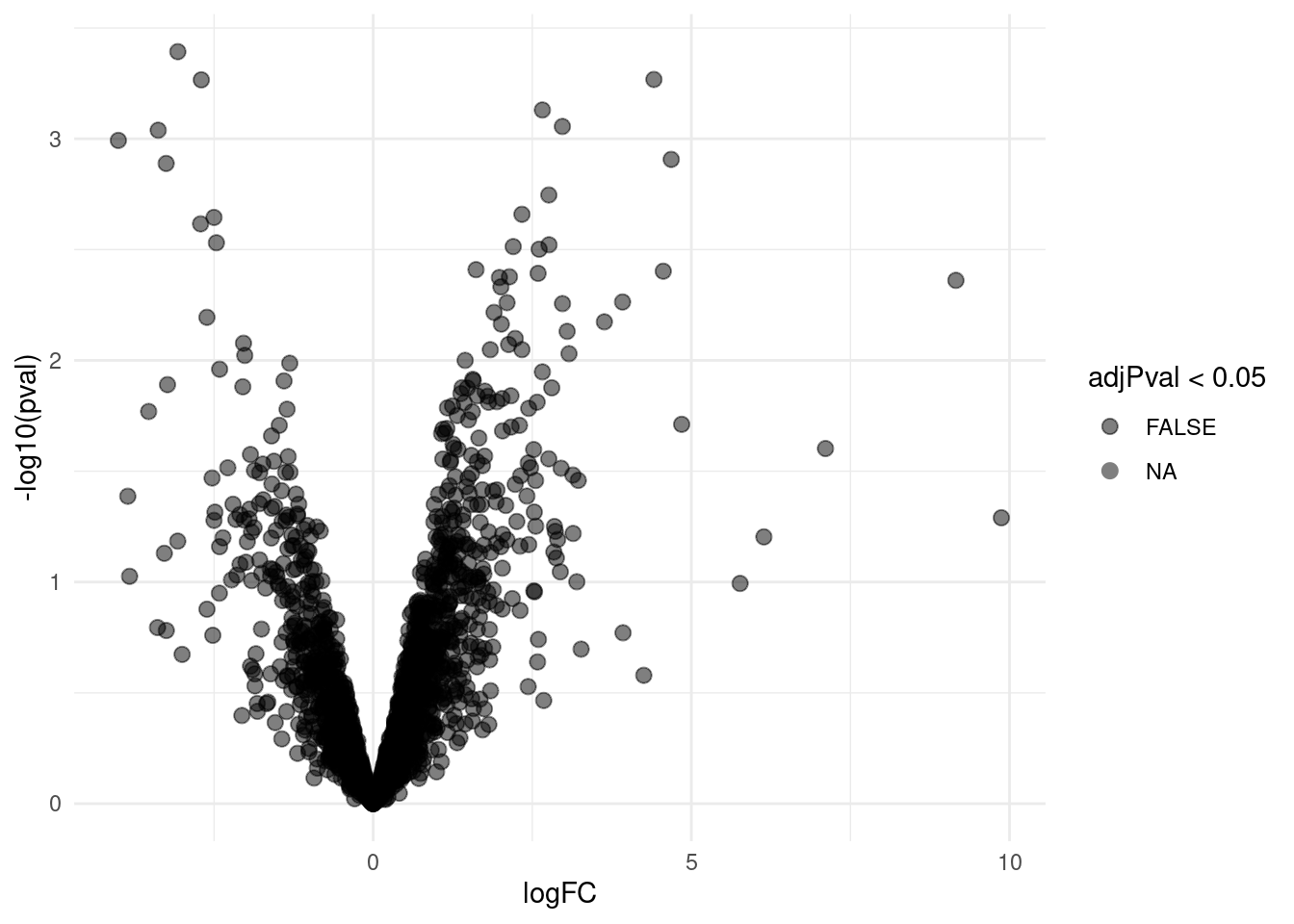
4.6.2 Heatmap
There were no genes significant at the 5% FDR level. We return the top 25 genes.
sigNamesInt <- rowData(pe[["proteinRobust"]])$"locationR:tissueV" %>%
rownames_to_column("proteinRobust") %>%
filter(adjPval<0.05) %>%
pull(proteinRobust)
hlp <- order((rowData(pe[["proteinRobust"]])$"locationR:tissueV")[,"adjPval"])[1:25]
heatmap(assay(pe[["proteinRobust"]])[hlp, ])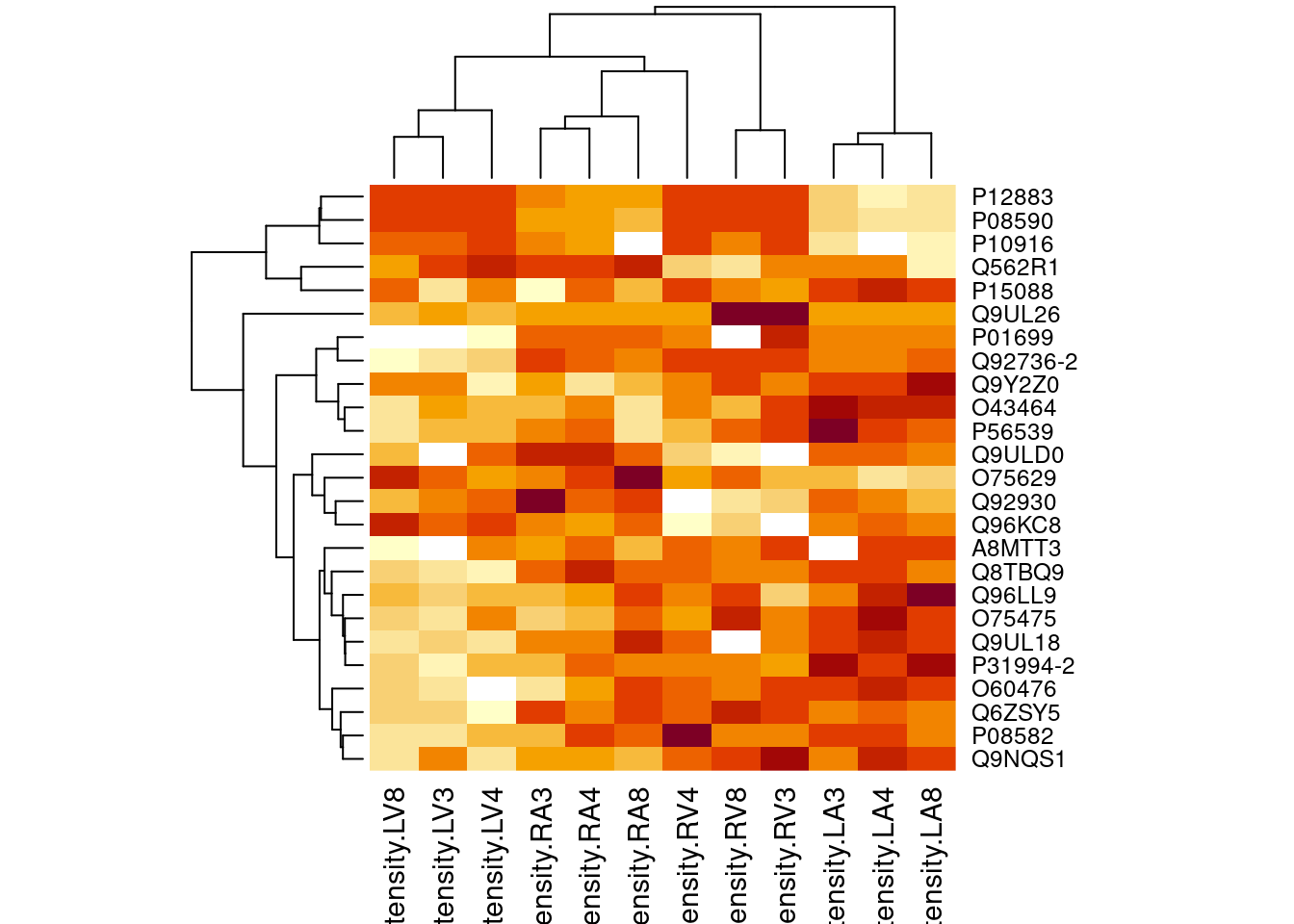
There are 0 proteins significantly differentially expressed at the 5% FDR level.
rowData(pe[["proteinRobust"]])$"locationR:tissueV" %>%
cbind(.,rowData(pe[["proteinRobust"]])$Protein.names) %>%
na.exclude %>%
filter(adjPval<0.05) %>%
arrange(pval)5 Large difference in number of proteins that are returned
Note, that much more proteins are returned significant for average contrast (\(\log_2 FC_{V-A}\)) as compared to contrast for assessing the fold change between ventriculum and atrium left and right. The power for the average contrast is larger than for the contrast left or right because the log2 FC can be estimated with higher precision.
For none of the proteins the interaction was significant (change in log2 FC between ventriculum and atrium in the right vs the left heart region). The power for the interaction is typically low.
5.1 Reason
Part of variance covariance matrix of model parameters due to design:
Variance of contrasts (diagonal elements) due to design
## tissueV tissueV + locationR:tissueV
## 0.6666667 0.6666667
## tissueV + 0.5 * locationR:tissueV locationR:tissueV
## 0.3333333 1.3333333## tissueV tissueV + locationR:tissueV
## 0.8164966 0.8164966
## tissueV + 0.5 * locationR:tissueV locationR:tissueV
## 0.5773503 1.1547005So it is clear that the standard error of the log2 FC left and right is the same. That of the average contrast is a factor \(\sqrt{2}\) smaller! Indeed, we use double the number of samples to estimate it!
## tissueV + 0.5 * locationR:tissueV
## 0.5## tissueV + 0.5 * locationR:tissueV
## 0.7071068## [1] 0.7071068The standard error of the interaction is a factor \(\sqrt{2}\) larger than that of the main effects!
## locationR:tissueV
## 2## locationR:tissueV
## 1.414214## [1] 1.4142145.2 Msqrob
This is not the case for the standard errors of protein 2???
Because msqrob is using robust regression to assess DE!
pe %>%
colData %>%
as_tibble %>%
mutate(w=getModel(rowData(pe[["proteinRobust"]])$msqrobModels[[2]])$w)For protein 2 the samples at the left and right side have different weights!
5.2.1 Part of standard error due to design:
covUnscaledRobust <- solve(
t(X) %*%
diag(
getModel(rowData(pe[["proteinRobust"]])$msqrobModels[[2]])$w) %*% X)
varContrastsRobust <- t(L)%*%covUnscaledRobust%*%L %>%
diag
varContrastsRobust## tissueV tissueV + locationR:tissueV
## 0.7161526 0.8673176
## tissueV + 0.5 * locationR:tissueV locationR:tissueV
## 0.3937862 1.5917957## tissueV tissueV + locationR:tissueV
## 0.8462580 0.9312989
## tissueV + 0.5 * locationR:tissueV locationR:tissueV
## 0.6275239 1.26166395.2.2 Standard errors Contrasts
## tissueV tissueV + locationR:tissueV
## 0.5569921 0.6129645
## tissueV + 0.5 * locationR:tissueV locationR:tissueV
## 0.4130252 0.8304050