Course Description
Mass spectrometry based proteomic experiments generate ever larger datasets and, as a consequence, complex data interpretation challenges. This course focuses on the statistical concepts for peptide identification, quantification, and differential analysis. Moreover, more advanced experimental designs and blocking will also be introduced. The core focus will be on shotgun proteomics data, and quantification using label-free precursor peptide (MS1) ion intensities. The course will rely exclusively on free and userfriendly opensource tools in R/Bioconductor. The course will provide a solid basis for beginners, but will also bring new perspectives to those already familiar with standard data interpretation procedures in proteomics.
Students can sharpen their background knowledge on Mass Spectrometry, Proteomics & Bioinformatics for Proteomics here: Mass Spectrometry and Bioinformatics for Proteomics
Target Audience
This course is oriented towards biologists and bioinformaticians with a particular interest in differential analysis for quantitative proteomics.
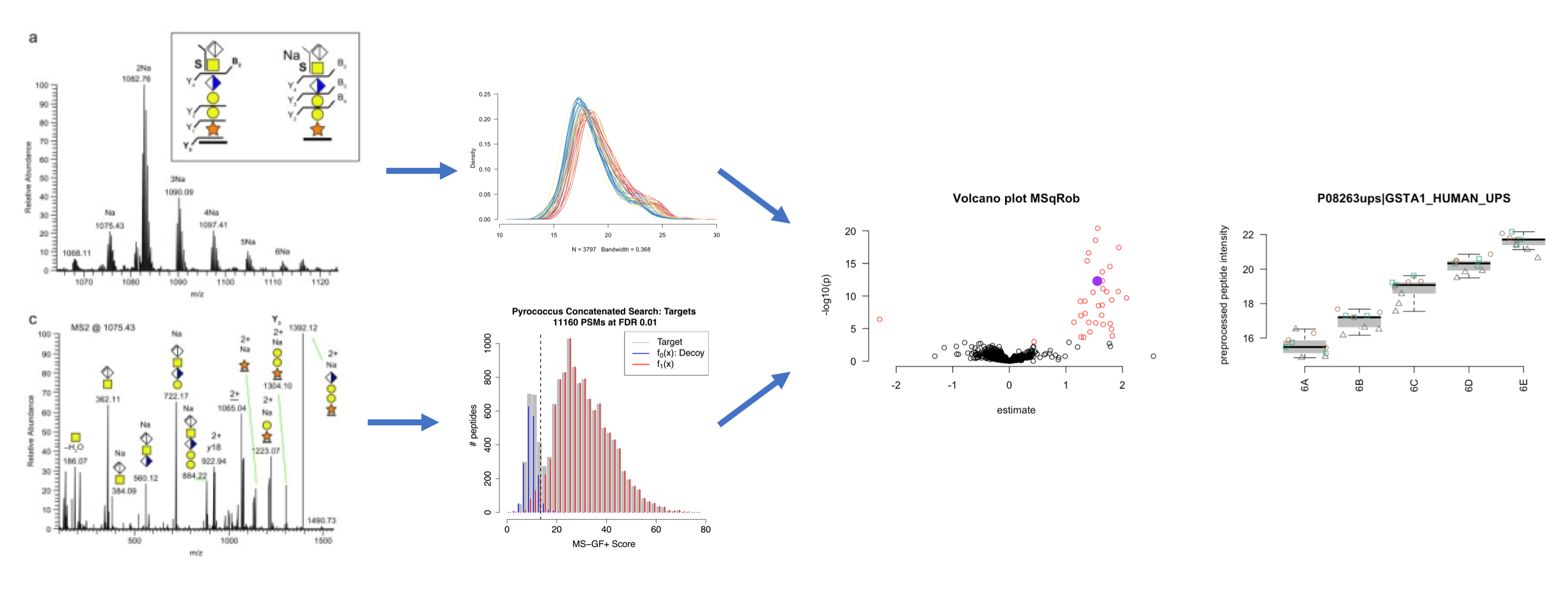
Statistical Data Analysis for Proteomics
Software and Data
Detailed Program
- Identification
- Preprocessing & Analysis of Label Free Quantitative Proteomics Experiments with Simple Designs
- Slides: Preprocessing, Results of Preprocessing
- Tutorial: Preprocessing & Analysis with MSqRob for Simple Designs
- Robust Regression: Robust Regression Explained by Example
- Statistical Inference & Analysis of Experiments with Factorial Designs